A head-to-toe dimerization has physiological relevance for ligand-induced inactivation of protein tyrosine receptor type Z.
Fujikawa, A., Sugawara, H., Tanga, N., Ishii, K., Kuboyama, K., Uchiyama, S., Suzuki, R., Noda, M.(2019) J Biological Chem 294: 14953-14965
- PubMed: 31416834 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.007878
- Primary Citation Related Structures:
6J6U - PubMed Abstract:
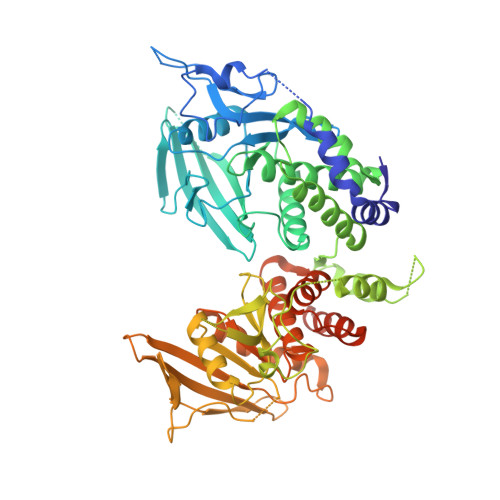
Protein-tyrosine phosphatase (PTPase) receptor type Z (PTPRZ) has two receptor isoforms, PTPRZ-A and -B, containing tandem intracellular PTP-D1 and -D2 domains, with only D1 being active. Pleiotrophin (PTN) binding to the extracellular PTPRZ region leads to inactivation of its PTPase activity, thereby facilitating oligodendrocyte precursor cell (OPC) differentiation and myelination in the central nervous system. However, the mechanisms responsible for PTN-induced PTPRZ inactivation remain unclear. We herein report that the crystal structure of the intracellular region of PTPRZ (PTPRZ-ICR) shows a "head-to-toe"-type dimer conformation, with D2 masking the catalytic site of D1. MS analyses revealed that PTPRZ-ICR proteins remain in monomer-dimer equilibrium in aqueous solution and that a substrate-derived inhibitory peptide or competitive inhibitor (SCB4380) specifically bind to the monomer form in a 1:1 ratio. A D2 deletion (ΔD2) or dimer interface mutation (DDKK) disrupted dimer formation, but SCB4380 binding was maintained. Similar to WT PTPRZ-B, monomer-biased PTPRZ-B-ΔD2 and PTPRZ-B-DDKK variants efficiently dephosphorylated p190RhoGAP at Tyr-1105 when co-expressed in BHK-21 cells. The catalytic activities of these variants were not suppressed by PTN treatment, but were inhibited by the cell-permeable PTPase inhibitor NAZ2329. Of note, the PTN treatment did not enhance OPC differentiation in primary cultured glial cells from ΔD2 or PTPase-inactive PTPRZ-B (CS) mutant knock-in mice. Our results thus indicate that PTN-induced PTPRZ inactivation results from dimer formation of the intracellular tandem PTP domains in a head-to-toe configuration, which is physiologically relevant to the control of OPC differentiation in vivo .
- Division of Molecular Neurobiology, National Institute for Basic Biology (NIBB), 5-1 Higashiyama, Myodaiji-cho, Okazaki, Aichi 444-8787, Japan.
Organizational Affiliation: