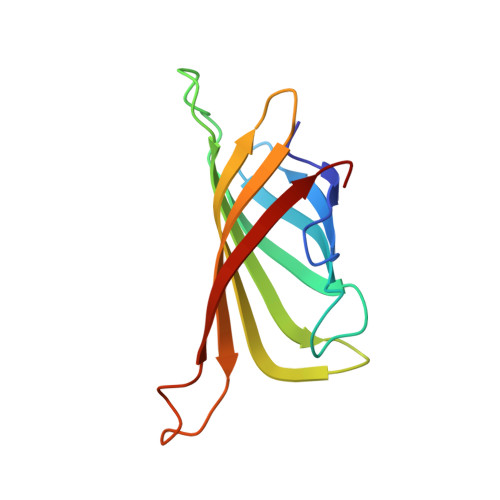
Single particle cryo-EM reconstruction of 52 kDa streptavidin at 3.2 Angstrom resolution.
Fan, X., Wang, J., Zhang, X., Yang, Z., Zhang, J.C., Zhao, L., Peng, H.L., Lei, J., Wang, H.W.(2019) Nat Commun 10: 2386-2386
- PubMed: 31160591 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-10368-w
- Primary Citation Related Structures:
6J6J, 6J6K - PubMed Abstract:
The fast development of single-particle cryogenic electron microscopy (cryo-EM) has made it more feasible to obtain the 3D structure of well-behaved macromolecules with a molecular weight higher than 300 kDa at ~3 Å resolution. However, it remains a challenge to obtain the high-resolution structures of molecules smaller than 200 kDa using single-particle cryo-EM. In this work, we apply the Cs-corrector-VPP-coupled cryo-EM to study the 52 kDa streptavidin (SA) protein supported on a thin layer of graphene and embedded in vitreous ice. We are able to solve both the apo-SA and biotin-bound SA structures at near-atomic resolution using single-particle cryo-EM. We demonstrate that the method has the potential to determine the structures of molecules as small as 39 kDa.
- Ministry of Education Key Laboratory of Protein Sciences, Beijing Advanced Innovation Center for Structural Biology, Beijing Frontier Research Center of Biological Structures, School of Life Sciences, Tsinghua University, Beijing, 100084, China.
Organizational Affiliation: