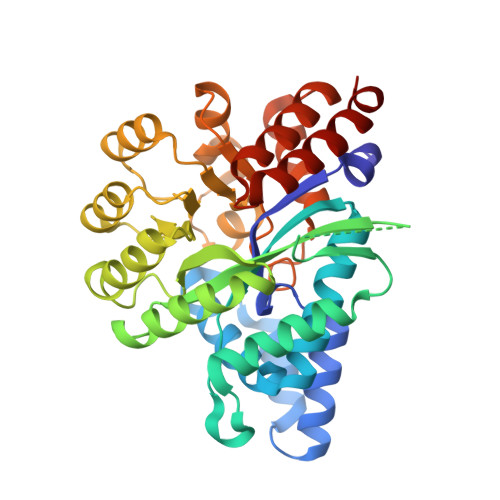
Structure ofArabidopsis thaliana N6-methyl-AMP deaminase ADAL with bound GMP and IMP and implications forN6-methyl-AMP recognition and processing.
Wu, B., Zhang, D., Nie, H., Shen, S., Li, Y., Li, S.(2019) RNA Biol 16: 1504-1512
- PubMed: 31318636
- DOI: https://doi.org/10.1080/15476286.2019.1642712
- Primary Citation Related Structures:
6IV5, 6J23, 6J4T - PubMed Abstract:
Arabidopsis thaliana aminohydrolase ( At ADAL) has been shown to be involved in the metabolism of N 6 -methyl-AMP, a proposed intermediate during m 6 A-modified RNA metabolism, which can be subsequently incorporated into newly synthesized RNA by Pol II. It has been proposed that At ADAL will prevent N 6 -methyl-AMP reuse and catabolize it to inosine monophosphate (IMP). Here, we have solved the crystal structures of At ADAL in the apo form and in complex with GMP and IMP in the presence of Zn 2+ . We have identified the substrate-binding pocket of At ADAL and compared it with that for adenosine deaminase (ADA), adenine deaminase (ADE) and AMP deaminase (AMPD) from multiple species. The comparisons reveal that plant ADAL1 may have the potential ability to catalyze different alkyl-group substituted substrates.
- Department of Biology, Southern University of Science and Technology , Shenzhen , Guangdong , China.
Organizational Affiliation: