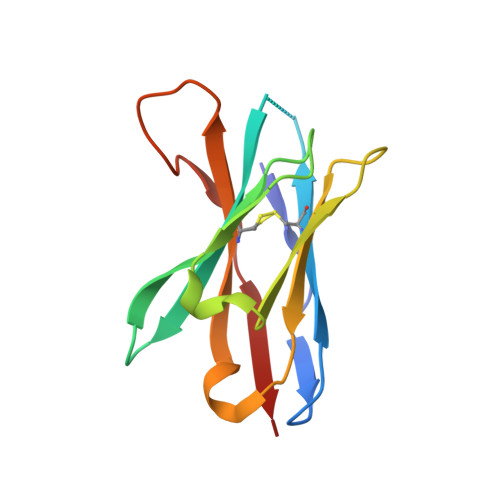
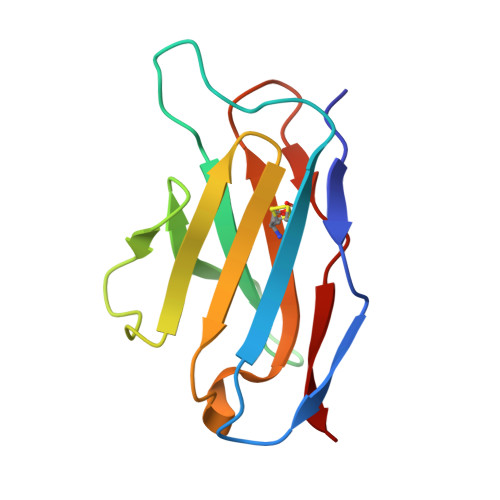
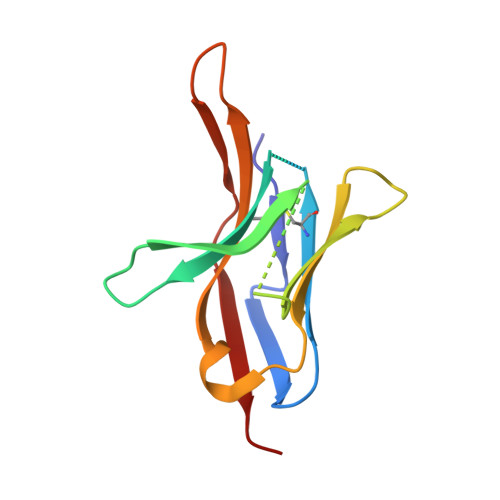
The FG Loop of PD-1 Serves as a "Hotspot" for Therapeutic Monoclonal Antibodies in Tumor Immune Checkpoint Therapy.
Chen, D., Tan, S., Zhang, H., Wang, H., He, W., Shi, R., Tong, Z., Zhu, J., Cheng, H., Gao, S., Chai, Y., Qi, J., Xiao, M., Yan, J., Gao, G.F.(2019) iScience 14: 113-124
- PubMed: 30952089 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.isci.2019.03.017
- Primary Citation Related Structures:
6J14, 6J15 - PubMed Abstract:
Programmed cell death 1 (PD-1)/PD-1 ligand-1 (PD-L1)-blocking monoclonal antibodies (mAbs) have taken center stage for tumor immune checkpoint therapy. Identification of the "hotspots" on PD-1 for mAbs will help to develop next-generation oral deliverable agents with long-lasting efficacy. Here, we identified two PD-1-targeting mAbs, GY-5 and GY-14, with PD-1/PD-L1-blocking efficacy. Complex structural information revealed that both mAbs mainly bind to the FG loop of PD-1, which also contributes multiple interactions with PD-L1. The FG loop adopts substantially varied conformations upon binding to different mAbs, providing a novel targetable region for the development of PD-1-specific biologics and small chemical molecules. Glycosylation modifications of PD-1 could be observed in three of the four potential N-linked glycosylation sites. However, the binding of GY-5 and GY-14 to PD-1 was not affected by glycosylation. These findings broaden our understanding of the mechanism of anti-PD-1 mAbs and provide insight into the development of agents targeting PD-1.
- CAS Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, Beijing 100101, China.
Organizational Affiliation: