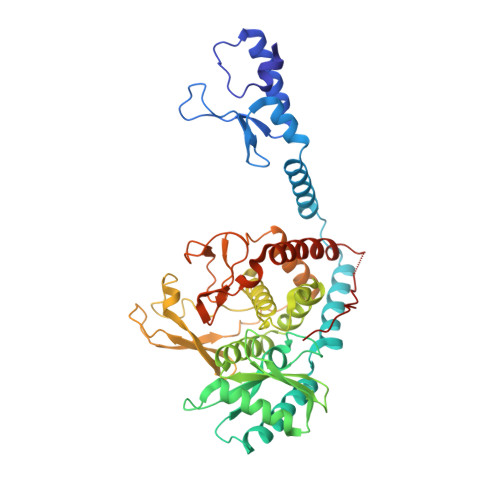
Crystal structure of the Lin28-interacting module of human terminal uridylyltransferase that regulates let-7 expression.
Yamashita, S., Nagaike, T., Tomita, K.(2019) Nat Commun 10: 1960-1960
- PubMed: 31036859 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-09966-5
- Primary Citation Related Structures:
6IW6 - PubMed Abstract:
Lin28-dependent oligo-uridylylation of precursor let-7 (pre-let-7) by terminal uridylyltransferase 4/7 (TUT4/7) represses let-7 expression by blocking Dicer processing, and regulates cell differentiation and proliferation. The interaction between the Lin28:pre-let-7 complex and the N-terminal Lin28-interacting module (LIM) of TUT4/7 is required for pre-let-7 oligo-uridylylation by the C-terminal catalytic module (CM) of TUT4/7. Here, we report crystallographic and biochemical analyses of the LIM of human TUT4. The LIM consists of the N-terminal Cys2His2-type zinc finger (ZF) and the non-catalytic nucleotidyltransferase domain (nc-NTD). The ZF of LIM adopts a distinct structural domain, and its structure is homologous to those of double-stranded RNA binding zinc fingers. The interaction between the ZF and pre-let-7 stabilizes the Lin28:pre-let-7:TUT4 ternary complex, and enhances the oligo-uridylylation reaction by the CM. Thus, the ZF in LIM and the zinc-knuckle in the CM, which interacts with the oligo-uridylylated tail, together facilitate Lin28-dependent pre-let-7 oligo-uridylylation.
- Department of Computational Biology and Medical Sciences, Graduate School of Frontier Sciences, The University of Tokyo, Kashiwa, Chiba, 277-8562, Japan.
Organizational Affiliation: