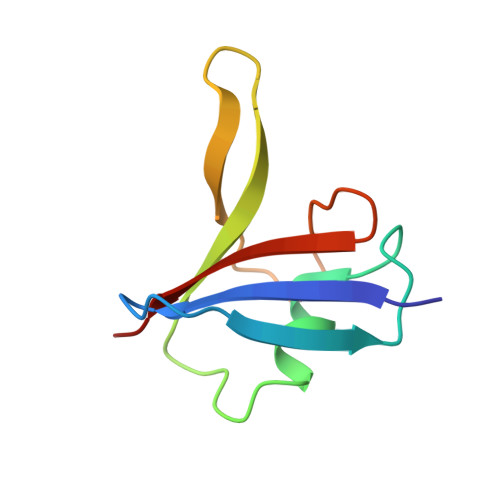
High-resolution structure of a Y27W mutant of the Dishevelled2 DIX domain.
Yamanishi, K., Sin, Y., Terawaki, S.I., Higuchi, Y., Shibata, N.(2019) Acta Crystallogr F Struct Biol Commun 75: 116-122
- PubMed: 30713163 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X18018290
- Primary Citation Related Structures:
6IW3 - PubMed Abstract:
Dishevelled (Dvl) is a positive regulator of the canonical Wnt pathway that downregulates the phosphorylation of β-catenin and its subsequent degradation. Dvl contains an N-terminal DIX domain, which is involved in its homooligomerization and interactions with regulators of the Wnt pathway. The crystal structure of a Y27W mutant of the Dishevelled2 DIX domain (DIX-Y27W) has been determined at 1.64 Å resolution. DIX-Y27W has a compact ubiquitin-like fold and self-associates with neighbouring molecules through β-bridges, resulting in a head-to-tail helical molecular arrangement similar to previously reported structures of DIX domains. Glu23 of DIX-Y27W forms a hydrogen bond to the side chain of Trp27, corresponding to the Glu762...Trp766 hydrogen bond of the rat Axin DIX domain, whereas Glu23 in the Y27D mutant of the Dishevelled2 DIX domain forms a salt bridge to Lys68 of the adjacent molecule. The high-resolution DIX-Y27W structure provides details of the head-to-tail interaction, including solvent molecules, and also the plausibly wild-type-like structure of the self-association surface compared with previously published Dvl DIX-domain mutants.
- Graduate School of Life Science, University of Hyogo, 3-2-1 Koto, Kamigori, Ako-gun, Hyogo 678-1297, Japan.
Organizational Affiliation: