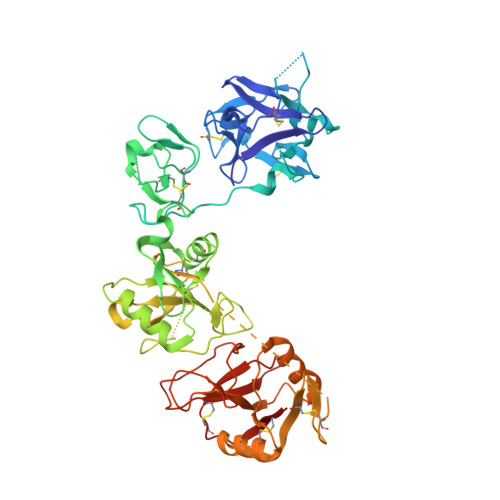
Structural basis of the pH-dependent conformational change of the N-terminal region of human mannose receptor/CD206.
Hu, Z., Wang, Y., Cheng, C., He, Y.(2019) J Struct Biol 208: 107384-107384
- PubMed: 31491467 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2019.09.001
- Primary Citation Related Structures:
6INN, 6INO, 6INU, 6INV, 6IOE - PubMed Abstract:
Mannose receptor (MR, CD206) is an immune receptor highly expressed on macrophages and plays important roles in glycoprotein clearance, immune response and matrix turnover. Previous studies have shown that MR recognizes multiple ligands and recycles between cell surface and endosomes, and the conformation and ligand binding of MR are regulated by environmental pH. However, due to the lack of high-resolution details, the mechanisms of the pH-dependent properties of MR have not been fully understood. Here we investigate the pH-dependent conformational change of MR by solving a series of crystal structures of MR N-terminal fragments (CysR~CTLD2/3) at pH ranging from 4.0 to 8.5. The results show that the CTLD3 domain plays a critical role in regulating the conformational change of the N-terminal region of MR by forming interactions with the CTLD2 domain specifically at acidic pH. Moreover, the structural data also show the conformational changes of the 4-SO 4 -GalNAc binding pocket at the CysR domain, which might be relevant to the binding and release of the ligand. Overall, these results provide a model for the pH-dependent conformational change of the N-terminal region of MR that may help to understand its functional mechanism at molecular level.
- National Center for Protein Science Shanghai, Shanghai Science Research Center; CAS Center for Excellence in Molecular Cell Science, Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences; University of Chinese Academy of Sciences, Shanghai, China.
Organizational Affiliation: