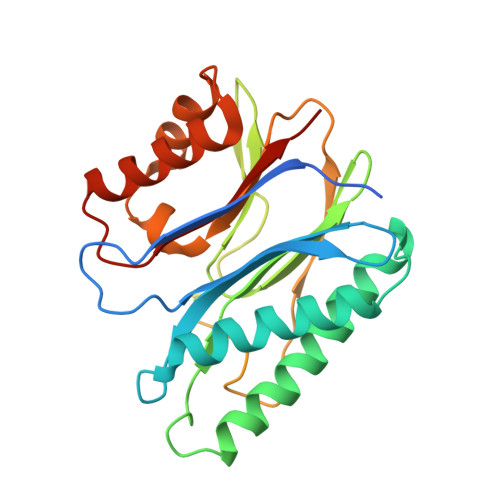
Structural Insight into the Mechanism of Staphylococcus aureus Stp1 Phosphatase.
Yang, T., Liu, T., Gan, J., Yu, K., Chen, K., Xue, W., Lan, L., Yang, S., Yang, C.G.(2019) ACS Infect Dis 5: 841-850
- PubMed: 30868877 Search on PubMed
- DOI: https://doi.org/10.1021/acsinfecdis.8b00316
- Primary Citation Related Structures:
6IHL, 6IHR, 6IHS, 6IHT, 6IHU, 6IHV, 6IHW - PubMed Abstract:
Staphylococcus aureus Stp1, which belongs to the bacterial metal-dependent protein phosphatase (PPM) family, is a promising candidate for antivirulence targeting. How Stp1 recognizes the phosphorylated peptide remains unclear, however. In order to investigate the recognition mechanism of Stp1 in depth, we have determined a series of crystal structures of S. aureus Stp1 in different states and the structural complex of Stp1 bound with a phosphorylated peptide His12. Different phosphorylated peptides, including MgrA- and GraR-derived phosphopeptides, are substrates of Stp1, which supports the function of Stp1 as a selective Ser/Thr phosphatase. In addition, interestingly, the crystal structures of R161-Stp1 variants combined with the biochemical activity validations have uncovered that R161 residue plays a key role to control the conformation switches of the flap domain in order to facilitate substrate binding and the dephosphorylation process. Our findings provide crucial structural insight into the molecular mechanism of S. aureus Stp1 phosphatase and reveal the phosphorylated peptides for biochemistry study and inhibitor screening of Stp1.
- State Key Laboratory Breeding Base of Green Pesticide and Agricultural Bioengineering, Key Laboratory of Green Pesticide and Agricultural Bioengineering, Ministry of Education, Center for R&D of Fine Chemicals , Guizhou University , 2708 South Huaxi Road , Guiyang , Guizhou 550025 , P. R. China.
Organizational Affiliation: