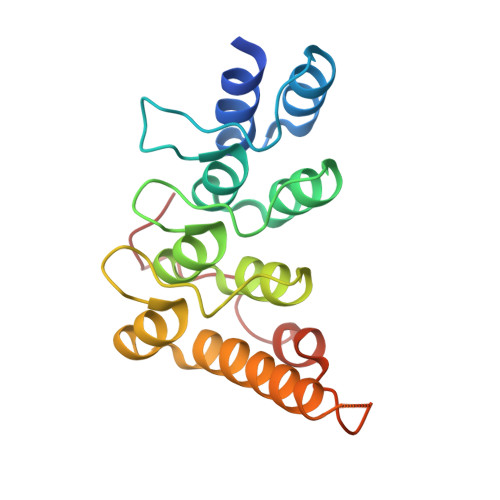
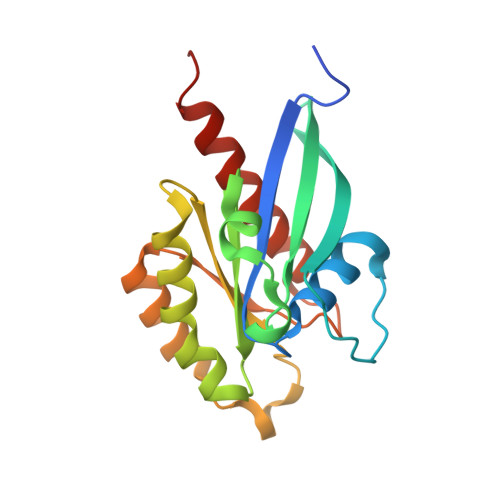
Rab35/ACAP2 and Rab35/RUSC2 Complex Structures Reveal Molecular Basis for Effector Recognition by Rab35 GTPase.
Lin, L., Shi, Y., Wang, M., Wang, C., Zhu, J., Zhang, R.(2019) Structure 27: 729
- PubMed: 30905672
- DOI: https://doi.org/10.1016/j.str.2019.02.008
- Primary Citation of Related Structures:
6IF2, 6IF3 - PubMed Abstract:
Rab35, a master regulator of membrane trafficking, regulates diverse cellular processes and is associated with various human diseases. Although a number of effectors have been identified, the molecular basis of Rab35-effector interactions remains unclear. Here, we provide the high-resolution crystal structures of Rab35 in complex with its two specific effectors ACAP2 and RUSC2, respectively. In the Rab35/ACAP2 complex structure, Rab35 binds to the terminal ankyrin repeat and a C-terminal extended α helix of ACAP2, revealing a previously uncharacterized binding mode both for Rabs and ankyrin repeats. In the Rab35/RUSC2 complex structure, Arg1015 of RUSC2 functions as a "pseudo-arginine finger" that stabilizes the GTP-bound Rab35, thus facilitating the assembly of Rab35/RUSC2 complex. The structural analysis allows us to design specific Rab35 mutants capable of eliminating Rab35/ACAP2 and Rab35/RUSC2 interactions, but not interfering with other effector bindings. The atomic structures also offer possible explanations to disease-associated mutants identified at the Rab35-effector interfaces.
- State Key Laboratory of Molecular Biology, CAS Center for Excellence in Molecular Cell Science, Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences, Shanghai Science Research Center, 333 Haike Road, Shanghai 201210, China.
Organizational Affiliation: