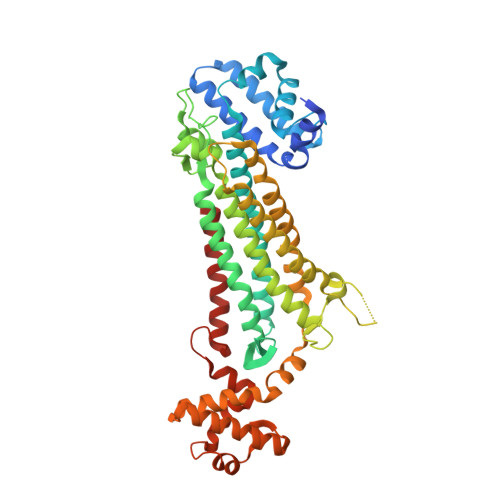
Structural studies on M. tuberculosis argininosuccinate lyase and its liganded complex: Insights into catalytic mechanism.
Paul, A., Mishra, A., Surolia, A., Vijayan, M.(2019) IUBMB Life 71: 643-652
- PubMed: 30615268 Search on PubMed
- DOI: https://doi.org/10.1002/iub.2000
- Primary Citation Related Structures:
6IEM, 6IEN - PubMed Abstract:
Argininosuccinate lyase catalyses the reversible breakdown of argininosuccinate into arginine and fumarate and is known to form tetramers in its quaternary association. The absence of structures involving competent enzymes bound to substrate/products came in the way of the precise elucidation of the catalytic mechanism of this family of proteins. Crystal structures of the enzyme from Mycobacterium tuberculosis in an unliganded form and its complex with the substrate/products have now been determined at 2.2 and 2.7 Å, respectively. The refinement of the structure of the complex was bedevilled by the presence of a lattice translocation defect. The two tetramers in the apo-crystals and the one in the crystals of the liganded protein, have the same structure except for the movements associated with enzyme action. Each molecule consists of an N-domain, an M-domain, and a C-domain. The molecule consists of four binding sites, each made up of peptide stretches from three subunits. Three binding sites appear to be occupied by the ligand in the transition state, while the products occupy the fourth site. The structure exhibits the movement of a loop in the M-domain and parts of the C-domain. This is the first instance when the appropriate movements are observed in a complex with bound substrate/product. The detailed picture of the binding site, active site residues and the movements associated with catalysis thus obtained, enabled a revisit of the mechanism of action of the enzyme. © 2019 IUBMB Life, 71(5):643-652, 2019.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore, India.
Organizational Affiliation: