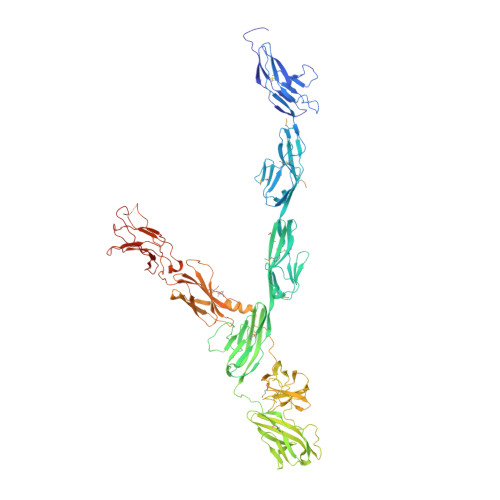
Structural Principles in Robo Activation and Auto-inhibition.
Barak, R., Yom-Tov, G., Guez-Haddad, J., Gasri-Plotnitsky, L., Maimon, R., Cohen-Berkman, M., McCarthy, A.A., Perlson, E., Henis-Korenblit, S., Isupov, M.N., Opatowsky, Y.(2019) Cell 177: 272-285.e16
- PubMed: 30853216 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2019.02.004
- Primary Citation Related Structures:
6I9S, 6IAA - PubMed Abstract:
Proper brain function requires high-precision neuronal expansion and wiring, processes controlled by the transmembrane Roundabout (Robo) receptor family and their Slit ligands. Despite their great importance, the molecular mechanism by which Robos' switch from "off" to "on" states remains unclear. Here, we report a 3.6 Å crystal structure of the intact human Robo2 ectodomain (domains D1-8). We demonstrate that Robo cis dimerization via D4 is conserved through hRobo1, 2, and 3 and the C. elegans homolog SAX-3 and is essential for SAX-3 function in vivo. The structure reveals two levels of auto-inhibition that prevent premature activation: (1) cis blocking of the D4 dimerization interface and (2) trans interactions between opposing Robo receptors that fasten the D4-blocked conformation. Complementary experiments in mouse primary neurons and C. elegans support the auto-inhibition model. These results suggest that Slit stimulation primarily drives the release of Robo auto-inhibition required for dimerization and activation.
- The Mina & Everard Goodman Faculty of Life Sciences, Bar-Ilan University, Israel.
Organizational Affiliation: