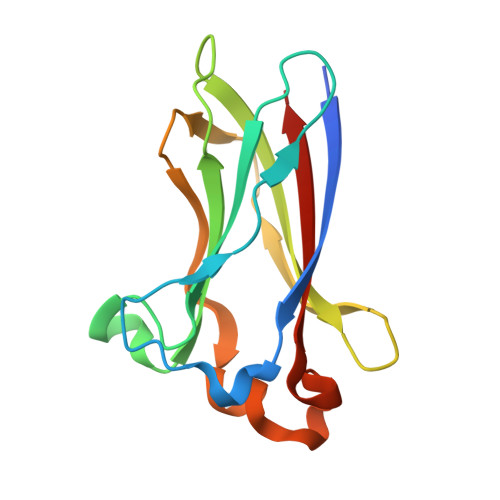
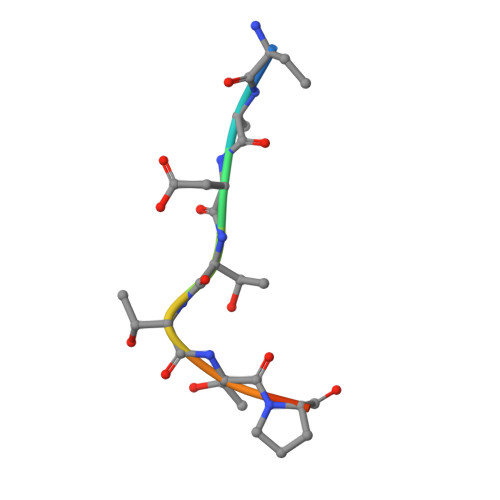
Structural Insights into BET Client Recognition of Endometrial and Prostate Cancer-Associated SPOP Mutants.
Ostertag, M.S., Hutwelker, W., Plettenburg, O., Sattler, M., Popowicz, G.M.(2019) J Mol Biology 431: 2213-2221
- PubMed: 31026449 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2019.04.017
- Primary Citation Related Structures:
6I41, 6I5P, 6I68, 6I7A - PubMed Abstract:
BET proteins such as BRD3 are oncogenic transcriptional coactivators. SPOP binding triggers their proteasomal degradation. In both endometrial and prostate cancers, SPOP mutations occur in the MATH domain, but with opposed influence on drug susceptibility. In prostate cancer, SPOP mutations presumably cause increased BET levels, decreasing BET inhibitor drug susceptibility. As opposed, in endometrial cancer, decreased BET levels concomitant with higher BET inhibitor drug susceptibility were observed. Here, we present the to our knowledge first co-crystal structure of SPOP and a bromodomain containing protein (BRD3). Our structural and biophysical data confirm the suggested loss-of-function in prostate cancer-associated SPOP mutants and provide mechanistic explanation. As opposed to previous literature, our data on endometrial cancer-associated SPOP mutants do not show altered binding behavior compared to wild-type SPOP, indicating a more complex regulatory mechanism. SPOP mutation screening may thus be considered a valuable personalized medicine tool for effective antitumor therapy.
- Institute of Structural Biology, Helmholtz Zentrum München, Ingolstädter Landstr. 1, 85764 Neuherberg, Germany; Center for Integrated Protein Science Munich at Chair of Biomolecular NMR Spectroscopy, Department Chemie, Technische Universität München, Lichtenbergstrasse 4, 85747 Garching, Germany.
Organizational Affiliation: