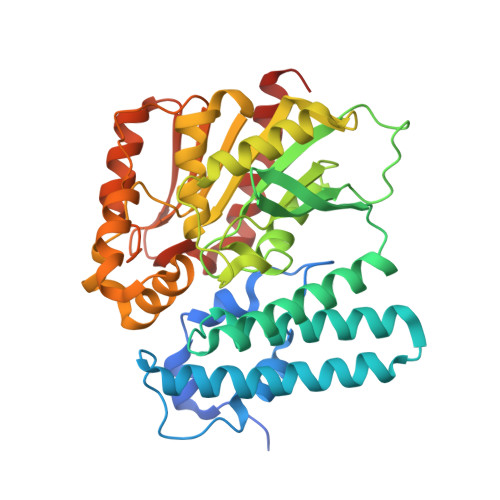
Molecular mechanism of polyketide shortening in anthraquinone biosynthesis ofPhotorhabdus luminescens.
Zhou, Q., Brauer, A., Adihou, H., Schmalhofer, M., Saura, P., Grammbitter, G.L.C., Kaila, V.R.I., Groll, M., Bode, H.B.(2019) Chem Sci 10: 6341-6349
- PubMed: 31341589 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/c9sc00749k
- Primary Citation Related Structures:
6HXA - PubMed Abstract:
Anthraquinones, a widely distributed class of aromatic natural products, are produced by a type II polyketide synthase system in the Gram-negative bacterium Photorhabdus luminescens . Heterologous expression of the antABCDEFGHI anthraquinone biosynthetic gene cluster in Escherichia coli identified AntI as an unusual lyase, catalysing terminal polyketide shortening prior to formation of the third aromatic ring. Functional in vitro and in vivo analysis of AntI using X-ray crystallography, structure-based mutagenesis, and molecular simulations revealed that AntI converts a defined octaketide to the tricyclic anthraquinone ring via retro-Claisen and Dieckmann reactions. Thus, AntI catalyses a so far unobserved multistep reaction in this PKS system.
- Molekulare Biotechnologie , Fachbereich Biowissenschaften , Buchmann Institute for Molecular Life Sciences (BMLS) , Goethe Universität Frankfurt , Max-von-Laue-Str. 15, Max-von-Laue-Str. 9 , 60438 Frankfurt am Main , Germany . Email: h.bode@bio.uni-frankfurt.de.
Organizational Affiliation: