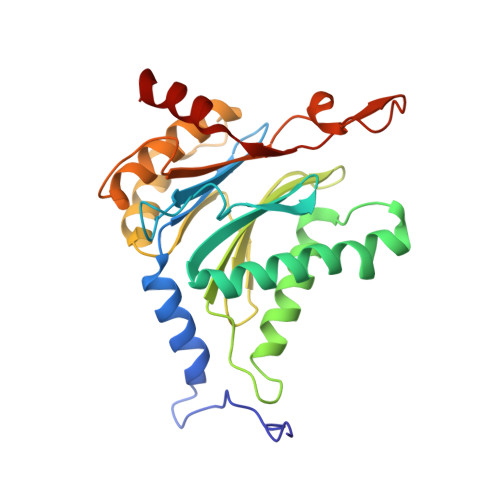
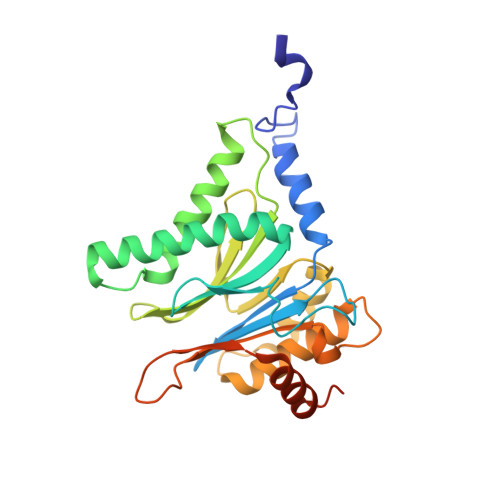
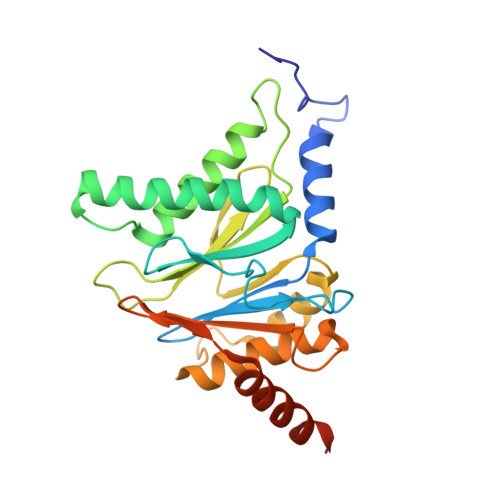
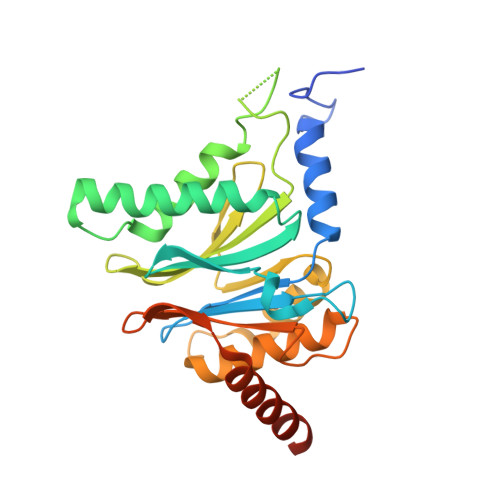
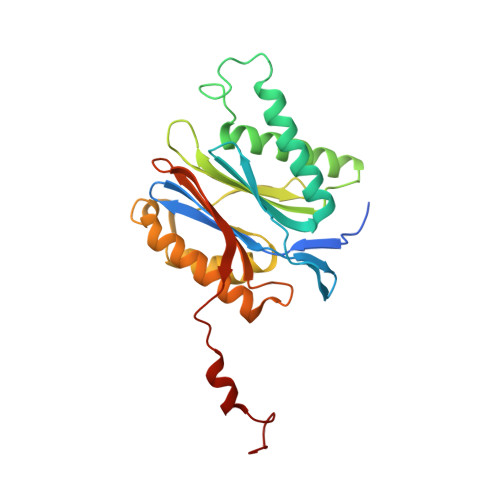
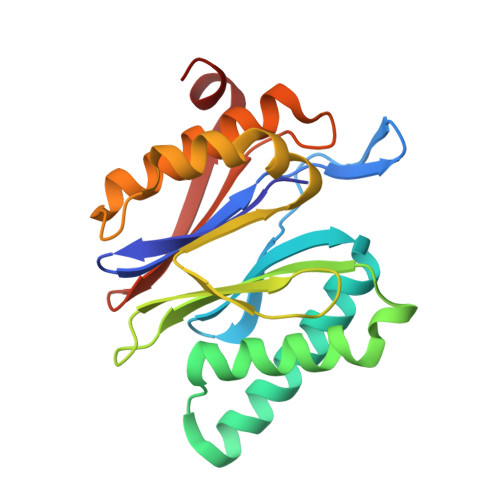
Structure-Based Design of Inhibitors Selective for Human Proteasome beta 2c or beta 2i Subunits.
Xin, B.T., Huber, E.M., de Bruin, G., Heinemeyer, W., Maurits, E., Espinal, C., Du, Y., Janssens, M., Weyburne, E.S., Kisselev, A.F., Florea, B.I., Driessen, C., van der Marel, G.A., Groll, M., Overkleeft, H.S.(2019) J Med Chem 62: 1626-1642
- PubMed: 30657666 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.8b01884
- Primary Citation Related Structures:
6HTB, 6HTC, 6HTD, 6HTP, 6HTR, 6HUB, 6HUC, 6HUQ, 6HUU, 6HUV, 6HV3, 6HV4, 6HV5, 6HV7, 6HVA, 6HVR, 6HVS, 6HVT, 6HVU, 6HVV, 6HVW, 6HVX, 6HVY, 6HW0, 6HW3, 6HW4, 6HW5, 6HW6, 6HW7, 6HW8, 6HW9, 6HWA, 6HWB, 6HWC, 6HWD, 6HWE, 6HWF - PubMed Abstract:
Subunit-selective proteasome inhibitors are valuable tools to assess the biological and medicinal relevance of individual proteasome active sites. Whereas the inhibitors for the β1c, β1i, β5c, and β5i subunits exploit the differences in the substrate-binding channels identified by X-ray crystallography, compounds selectively targeting β2c or β2i could not yet be rationally designed because of the high structural similarity of these two subunits. Here, we report the development, chemical synthesis, and biological screening of a compound library that led to the identification of the β2c- and β2i-selective compounds LU-002c (4; IC 50 β2c: 8 nM, IC 50 β2i/β2c: 40-fold) and LU-002i (5; IC 50 β2i: 220 nM, IC 50 β2c/β2i: 45-fold), respectively. Co-crystal structures with β2 humanized yeast proteasomes visualize protein-ligand interactions crucial for subunit specificity. Altogether, organic syntheses, activity-based protein profiling, yeast mutagenesis, and structural biology allowed us to decipher significant differences of β2 substrate-binding channels and to complete the set of subunit-selective proteasome inhibitors.
- Gorlaeus Laboratories , Leiden Institute of Chemistry and Netherlands Proteomics Centre , Einsteinweg 55 , 2333 CC Leiden , Netherlands.
Organizational Affiliation: