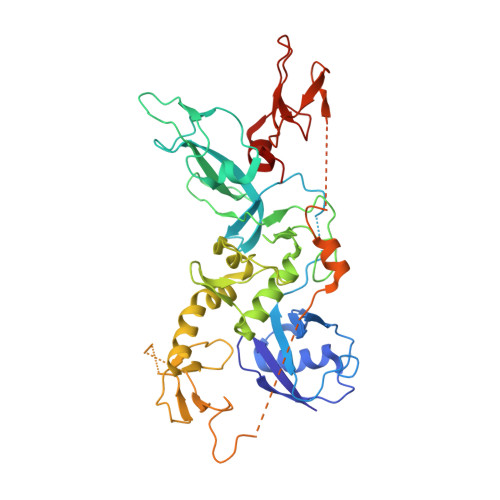
Phosphorylation of Parkin at serine 65 is essential for its activation in vivo .
McWilliams, T.G., Barini, E., Pohjolan-Pirhonen, R., Brooks, S.P., Singh, F., Burel, S., Balk, K., Kumar, A., Montava-Garriga, L., Prescott, A.R., Hassoun, S.M., Mouton-Liger, F., Ball, G., Hills, R., Knebel, A., Ulusoy, A., Di Monte, D.A., Tamjar, J., Antico, O., Fears, K., Smith, L., Brambilla, R., Palin, E., Valori, M., Eerola-Rautio, J., Tienari, P., Corti, O., Dunnett, S.B., Ganley, I.G., Suomalainen, A., Muqit, M.M.K.(2018) Open Biol 8
- PubMed: 30404819 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1098/rsob.180108
- Primary Citation Related Structures:
6HUE - PubMed Abstract:
Mutations in PINK1 and Parkin result in autosomal recessive Parkinson's disease (PD). Cell culture and in vitro studies have elaborated the PINK1-dependent regulation of Parkin and defined how this dyad orchestrates the elimination of damaged mitochondria via mitophagy. PINK1 phosphorylates ubiquitin at serine 65 (Ser65) and Parkin at an equivalent Ser65 residue located within its N-terminal ubiquitin-like domain, resulting in activation; however, the physiological significance of Parkin Ser65 phosphorylation in vivo in mammals remains unknown. To address this, we generated a Parkin Ser65Ala (S65A) knock-in mouse model. We observe endogenous Parkin Ser65 phosphorylation and activation in mature primary neurons following mitochondrial depolarization and reveal this is disrupted in Parkin S65A/S65A neurons. Phenotypically, Parkin S65A/S65A mice exhibit selective motor dysfunction in the absence of any overt neurodegeneration or alterations in nigrostriatal mitophagy. The clinical relevance of our findings is substantiated by the discovery of homozygous PARKIN ( PARK2 ) p.S65N mutations in two unrelated patients with PD. Moreover, biochemical and structural analysis demonstrates that the Parkin S65N/S65N mutant is pathogenic and cannot be activated by PINK1. Our findings highlight the central role of Parkin Ser65 phosphorylation in health and disease.
- MRC Protein Phosphorylation and Ubiquitylation Unit, School of Life Sciences, University of Dundee, Dundee DD1 5EH, UK thomas.mcwilliams@helsinki.fi.
Organizational Affiliation: