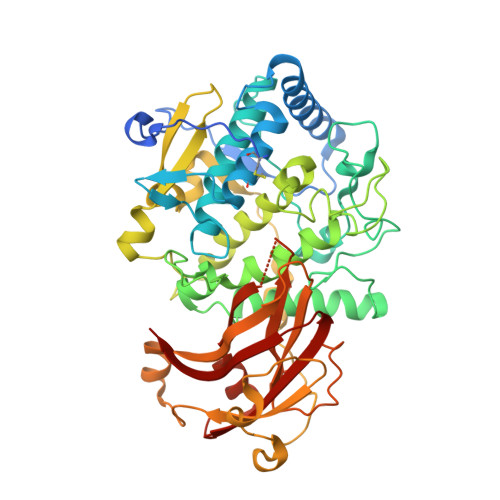
Biochemical and structural characterization of tomato polyphenol oxidases provide novel insights into their substrate specificity.
Kampatsikas, I., Bijelic, A., Rompel, A.(2019) Sci Rep 9: 4022-4022
- PubMed: 30858490 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-39687-0
- Primary Citation Related Structures:
6HQI, 6HQJ - PubMed Abstract:
Polyphenol oxidases (PPOs) contain the structurally similar enzymes tyrosinases (TYRs) and catechol oxidases (COs). Two cDNAs encoding pro-PPOs from tomato (Solanum lycopersicum) were cloned and heterologously expressed in Escherichia coli. The two pro-PPOs (SlPPO1-2) differ remarkably in their activity as SlPPO1 reacts with the monophenols tyramine (k cat = 7.94 s -1 ) and phloretin (k cat = 2.42 s -1 ) and was thus characterized as TYR, whereas SlPPO2 accepts only diphenolic substrates like dopamine (k cat = 1.99 s -1 ) and caffeic acid (k cat = 20.33 s -1 ) rendering this enzyme a CO. This study, for the first time, characterizes a plant TYR and CO originating from the same organism. Moreover, X-ray structure analysis of the latent holo- and apo-SlPPO1 (PDB: 6HQI and 6HQJ) reveals an unprecedented high flexibility of the gatekeeper residue phenylalanine (Phe270). Docking studies showed that depending on its orientation the gatekeeper residue could either stabilize and correctly position incoming substrates or hinder their entrance into the active site. Furthermore, phloretin, a substrate of SIPPO1 (K m = 0.11 mM), is able to approach the active centre of SlPPO1 with both phenolic rings. Kinetic and structural results indicate that phloretin could act as a natural substrate and connote the participation of PPOs in flavonoid-biosynthesis.
- Universität Wien, Fakultät für Chemie, Institut für Biophysikalische Chemie, Wien, Austria.
Organizational Affiliation: