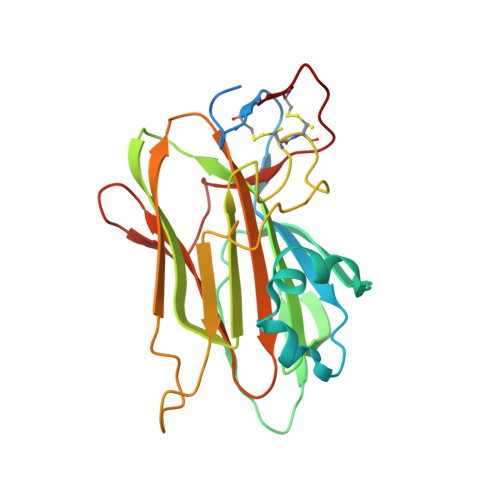
Structural and Functional Characterization of PA14/Flo5-Like Adhesins FromKomagataella pastoris.
Kock, M., Bruckner, S., Wozniak, N., Maestre-Reyna, M., Veelders, M., Schlereth, J., Mosch, H.U., Essen, L.O.(2018) Front Microbiol 9: 2581-2581
- PubMed: 30425696 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3389/fmicb.2018.02581
- Primary Citation Related Structures:
6HOS - PubMed Abstract:
Cell-cell and cell-substrate based adhesion of yeasts are major determinants of their adoption of different life styles. Genome-mining of ascomycetous GPI-anchored cell wall proteins with lectin-like PA14 domains identified a unique class of putative adhesins in the clade of methylotrophic Komagataella yeasts, many of which are known to colonize plants and insects involving yet unknown adhesion mechanisms. Here, we report the functional and structural analysis of two of its members: Kp Flo1 (=Cea1), that is highly specific for terminal N -acetylglucosamine moieties, and Kp Flo2, which represents an orphan lectin with intact binding site but unknown specificity. Crystal structures of the Cea1 adhesion domain complexed to N -acetylglucosamine and N , N '-diacetylchitobiose reveal a Ca 2+ -dependent binding mode that differs from other members of the PA14/Flo5 adhesin family. Heterologous expression of Cea1A in Saccharomyces cerevisiae promotes cellular adhesion to non-reducing ends of non-crystalline chitin. Overall, our data suggest that high-affinity recognition of β-GlcNAc-capped glycans by Cea1 enable Komagataella species to interact with surface cues present in fungi and insects.
- Department of Biochemistry, Faculty of Chemistry, Philipps University of Marburg, Marburg, Germany.
Organizational Affiliation: