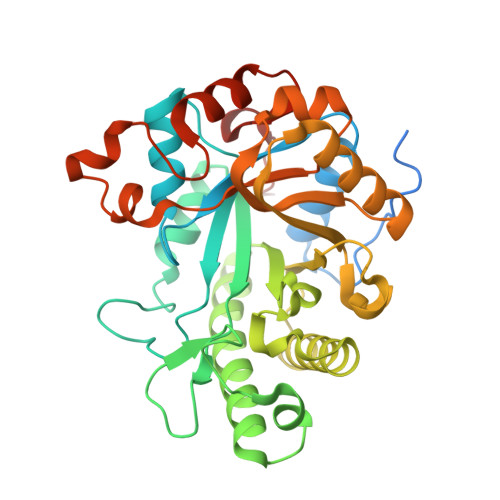
A surface-exposed GH26 beta-mannanase fromBacteroides ovatus: Structure, role, and phylogenetic analysis ofBoMan26B.
Bagenholm, V., Wiemann, M., Reddy, S.K., Bhattacharya, A., Rosengren, A., Logan, D.T., Stalbrand, H.(2019) J Biological Chem 294: 9100-9117
- PubMed: 31000630 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA118.007171
- Primary Citation Related Structures:
6HF2, 6HF4 - PubMed Abstract:
The galactomannan utilization locus ( Bo ManPUL) of the human gut bacterium Bacteroides ovatus encodes Bo Man26B, a cell-surface-exposed endomannanase whose functional and structural features have been unclear. Our study now places Bo Man26B in context with related enzymes and reveals the structural basis for its specificity. Bo Man26B prefers longer substrates and is less restricted by galactose side-groups than the mannanase Bo Man26A of the same locus. Using galactomannan, Bo Man26B generated a mixture of (galactosyl) manno-oligosaccharides shorter than mannohexaose. Three defined manno-oligosaccharides had affinity for the SusD-like surface-exposed glycan-binding protein, predicted to be implicated in saccharide transport. Co-incubation of Bo Man26B and the periplasmic α-galactosidase Bo Gal36A increased the rate of galactose release by about 10-fold compared with the rate without Bo Man26B. The results suggested that Bo Man26B performs the initial attack on galactomannan, generating oligosaccharides that after transport to the periplasm are processed by Bo Gal36A. A crystal structure of Bo Man26B with galactosyl-mannotetraose bound in subsites -5 to -2 revealed an open and long active-site cleft with Trp-112 in subsite -5 concluded to be involved in mannosyl interaction. Moreover, Lys-149 in the -4 subsite interacted with the galactosyl side-group of the ligand. A phylogenetic tree consisting of GH26 enzymes revealed four strictly conserved GH26 residues and disclosed that Bo Man26A and Bo Man26B reside on two distinct phylogenetic branches (A and B). The three other branches contain lichenases, xylanases, or enzymes with unknown activities. Lys-149 is conserved in a narrow part of branch B, and Trp-112 is conserved in a wider group within branch B.
- From the Department of Biochemistry and Structural Biology, Lund University P. O. Box 124, S-221 00, Lund, Sweden and.
Organizational Affiliation: