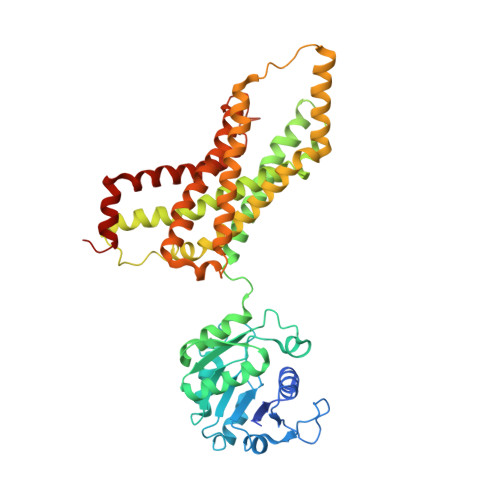
Cryo-EM structures of human STEAP4 reveal mechanism of iron(III) reduction.
Oosterheert, W., van Bezouwen, L.S., Rodenburg, R.N.P., Granneman, J., Forster, F., Mattevi, A., Gros, P.(2018) Nat Commun 9: 4337-4337
- PubMed: 30337524 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-06817-7
- Primary Citation Related Structures:
6HCY, 6HD1 - PubMed Abstract:
Enzymes of the six-transmembrane epithelial antigen of the prostate (STEAP) family reduce Fe 3+ and Cu 2+ ions to facilitate metal-ion uptake by mammalian cells. STEAPs are highly upregulated in several types of cancer, making them potential therapeutic targets. However, the structural basis for STEAP-catalyzed electron transfer through an array of cofactors to metals at the membrane luminal side remains elusive. Here, we report cryo-electron microscopy structures of human STEAP4 in absence and presence of Fe 3+ -NTA. Domain-swapped, trimeric STEAP4 orients NADPH bound to a cytosolic domain onto axially aligned flavin-adenine dinucleotide (FAD) and a single b-type heme that cross the transmembrane-domain to enable electron transfer. Substrate binding within a positively charged ring indicates that iron gets reduced while in complex with its chelator. These molecular principles of iron reduction provide a basis for exploring STEAPs as therapeutic targets.
- Crystal and Structural Chemistry, Bijvoet Center for Biomolecular Research, Department of Chemistry, Faculty of Science, Utrecht University, Padualaan 8, 3584 CH, Utrecht, The Netherlands.
Organizational Affiliation: