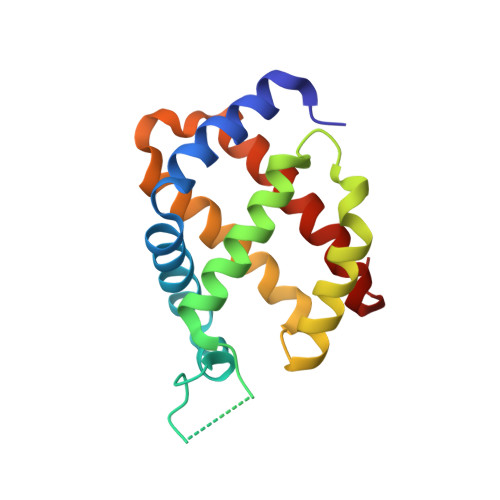
Proximal and distal control for ligand binding in neuroglobin: role of the CD loop and evidence for His64 gating.
Exertier, C., Milazzo, L., Freda, I., Montemiglio, L.C., Scaglione, A., Cerutti, G., Parisi, G., Anselmi, M., Smulevich, G., Savino, C., Vallone, B.(2019) Sci Rep 9: 5326-5326
- PubMed: 30926858 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-41780-3
- Primary Citation Related Structures:
6H5Z, 6H6C, 6H6I, 6H6J - PubMed Abstract:
Neuroglobin (Ngb) is predominantly expressed in neurons of the central and peripheral nervous systems and it clearly seems to be involved in neuroprotection. Engineering Ngb to observe structural and dynamic alterations associated with perturbation in ligand binding might reveal important structural determinants, and could shed light on key features related to its mechanism of action. Our results highlight the relevance of the CD loop and of Phe106 as distal and proximal controls involved in ligand binding in murine neuroglobin. We observed the effects of individual and combined mutations of the CD loop and Phe106 that conferred to Ngb higher CO binding velocities, which we correlate with the following structural observations: the mutant F106A shows, upon CO binding, a reduced heme sliding hindrance, with the heme present in a peculiar double conformation, whereas in the CD loop mutant "Gly-loop", the original network of interactions between the loop and the heme was abolished, enhancing binding via facilitated gating out of the distal His64. Finally, the double mutant, combining both mutations, showed a synergistic effect on CO binding rates. Resonance Raman spectroscopy and MD simulations support our findings on structural dynamics and heme interactions in wild type and mutated Ngbs.
- Dip. di Scienze Biochimiche "A. Rossi Fanelli", Sapienza Università di Roma, P.le A. Moro 5, 00185, Rome, Italy.
Organizational Affiliation: