Crystal Structure of protein MJ1004 from Mathanocaldococcus jannaschii
Alfonso Martinez-Cruz, L., Oyenarte, I., Fernandez-Rodriguez, C.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| MJ1004 | 214 | Methanocaldococcus jannaschii DSM 2661 | Mutation(s): 0 Gene Names: MJ1004 | 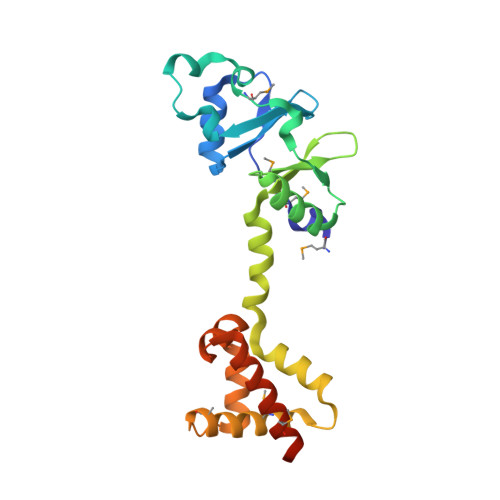 | |
UniProt | |||||
Find proteins for Q58410 (Methanocaldococcus jannaschii (strain ATCC 43067 / DSM 2661 / JAL-1 / JCM 10045 / NBRC 100440)) Explore Q58410 Go to UniProtKB: Q58410 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q58410 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MSE Query on MSE | A, B | L-PEPTIDE LINKING | C5 H11 N O2 Se |  | MET |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 80.899 | α = 90 |
| b = 80.899 | β = 90 |
| c = 178.95 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| Aimless | data scaling |
| PDB_EXTRACT | data extraction |
| AutoSol | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Spain | PI2010-17 | |
| Spain | ETORTEK IE05-147, IE07-202 | |
| Spain | Exp. 7/13/08/2006/11 | |
| Spain | Exp. 7/13/08/2005/14 | |
| Spain | MICINN BFU2010-17857 | |
| Spain | CSD2008-00005 |