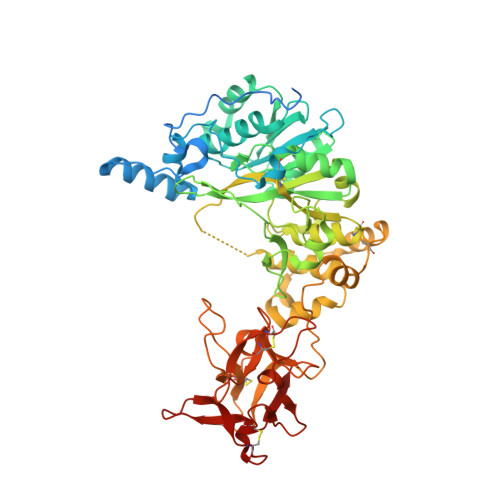
Structural and Mechanistic Insights into the Catalytic-Domain-Mediated Short-Range Glycosylation Preferences of GalNAc-T4.
de Las Rivas, M., Paul Daniel, E.J., Coelho, H., Lira-Navarrete, E., Raich, L., Companon, I., Diniz, A., Lagartera, L., Jimenez-Barbero, J., Clausen, H., Rovira, C., Marcelo, F., Corzana, F., Gerken, T.A., Hurtado-Guerrero, R.(2018) ACS Cent Sci 4: 1274-1290
- PubMed: 30276263 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acscentsci.8b00488
- Primary Citation Related Structures:
6H0B - PubMed Abstract:
Mucin-type O -glycosylation is initiated by a family of polypeptide GalNAc-transferases (GalNAc-Ts) which are type-II transmembrane proteins that contain Golgi luminal catalytic and lectin domains that are connected by a flexible linker. Several GalNAc-Ts, including GalNAc-T4, show both long-range and short-range prior glycosylation specificity, governed by their lectin and catalytic domains, respectively. While the mechanism of the lectin-domain-dependent glycosylation is well-known, the molecular basis for the catalytic-domain-dependent glycosylation of glycopeptides is unclear. Herein, we report the crystal structure of GalNAc-T4 bound to the diglycopeptide GAT*GAGAGAGT*TPGPG (containing two α-GalNAc glycosylated Thr (T*), the PXP motif and a "naked" Thr acceptor site) that describes its catalytic domain glycopeptide GalNAc binding site. Kinetic studies of wild-type and GalNAc binding site mutant enzymes show the lectin domain GalNAc binding activity dominates over the catalytic domain GalNAc binding activity and that these activities can be independently eliminated. Surprisingly, a flexible loop protruding from the lectin domain was found essential for the optimal activity of the catalytic domain. This work provides the first structural basis for the short-range glycosylation preferences of a GalNAc-T.
- BIFI, University of Zaragoza, BIFI-IQFR (CSIC) Joint Unit, Mariano Esquillor s/n, Campus Rio Ebro, Edificio I+D, Zaragoza 50018, Spain.
Organizational Affiliation: