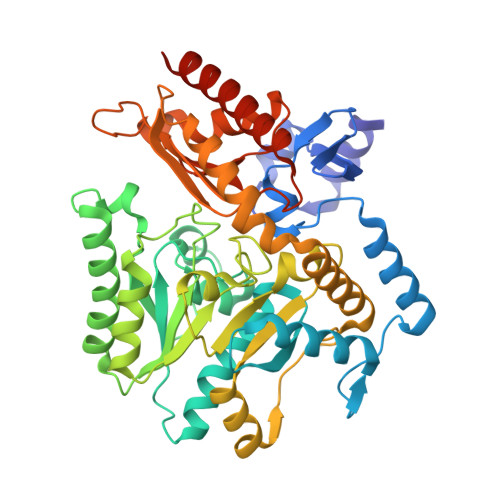
Widely applicable background depletion step enables transaminase evolution through solid-phase screening.
Planchestainer, M., Hegarty, E., Heckmann, C.M., Gourlay, L.J., Paradisi, F.(2019) Chem Sci 10: 5952-5958
- PubMed: 31360401 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/c8sc05712e
- Primary Citation Related Structures:
6GWI - PubMed Abstract:
Directed evolution of transaminases is a widespread technique in the development of highly sought-after biocatalysts for industrial applications. This process, however, is challenged by the limited availability of effective high-throughput protocols to evaluate mutant libraries. Here we report a rapid, reliable, and widely applicable background depletion method for solid-phase screening of transaminase variants, which was successfully applied to a transaminase from Halomonas elongata (HEWT), evolved through rounds of random mutagenesis towards a series of diverse prochiral ketones. This approach enabled the identification of transaminase variants in viable cells with significantly improved activity towards para-substituted acetophenones (up to 60-fold), as well as tetrahydrothiophen-3-one and related substrates. Rationalisation of the mutants was assisted by determination of the high-resolution wild-type HEWT crystal structure presented herein.
- University of Nottingham , School of Chemistry , University Park , Nottingham NG7 2RD , United Kingdom.
Organizational Affiliation: