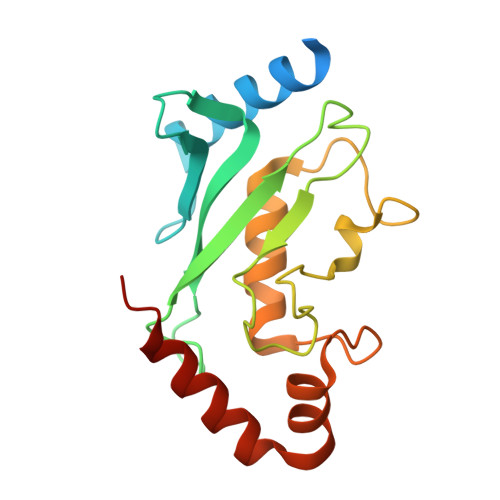
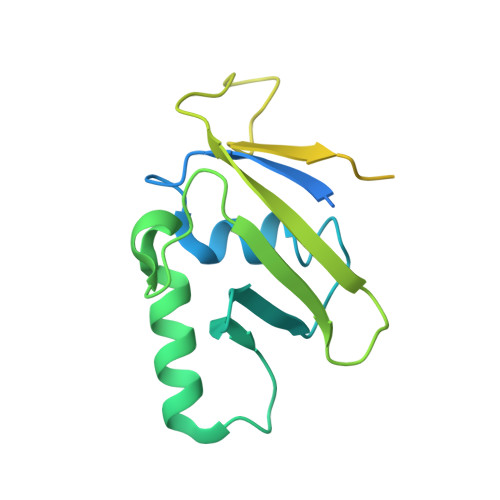
Structural insights into SUMO E1-E2 interactions in Arabidopsis uncovers a distinctive platform for securing SUMO conjugation specificity across evolution.
Liu, B., Lois, L.M., Reverter, D.(2019) Biochem J 476: 2127-2139
- PubMed: 31292170 Search on PubMed
- DOI: https://doi.org/10.1042/BCJ20190232
- Primary Citation Related Structures:
6GUM, 6GV3 - PubMed Abstract:
SUMOylation of proteins involves the concerted action of the E1-activating enzyme, E2-conjugating enzyme and E3-ligases. An essential discrimination step in the SUMOylation pathway corresponds to the initial interaction between E1 ubiquitin-fold domain (UFD) and E2 enzymes. Although E2 orthologs possess high sequence identity, the E2 binding region of the UFD domains has diverged across evolution. Moreover, in reciprocal in vitro conjugation reactions Arabidopsis E1 and E2 SCE1 fail to interact efficiently with cognate human E2 Ubc9 and E1 partners, respectively. To gain more insights into the properties of this interface in evolutionary distant organisms, we solved the crystal structure of SUMO E2 SCE1 and its complex with E1 UFD in Arabidopsis. In addition to a few common structural determinants, the interface between the E1 UFD and E2 in Arabidopsis is distinct compared with human and yeast, in particular by the presence of a longer α-helix in the Arabidopsis UFD domain. Despite the variability of E1 UFD domains in these surfaces, they establish specific interactions with highly conserved surfaces of their cognate E2 enzymes. Functional analysis of the different E2 interface residues between human and Arabidopsis revealed Val37 (Met36 in human), as a determinant that provides specificity in the E1-E2 recognition in plants.
- Departament de Bioquimica i Biologia Molecular, Institut de Biotecnologia i de Biomedicina, Serra Hunter Fellow, Universitat Autonoma de Barcelona, 08193 Barcelona, Bellaterra, Spain.
Organizational Affiliation: