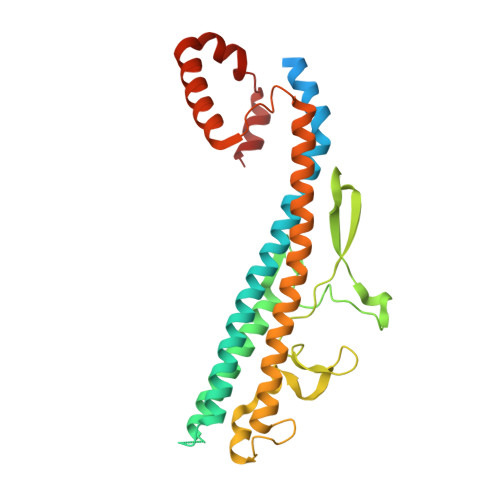
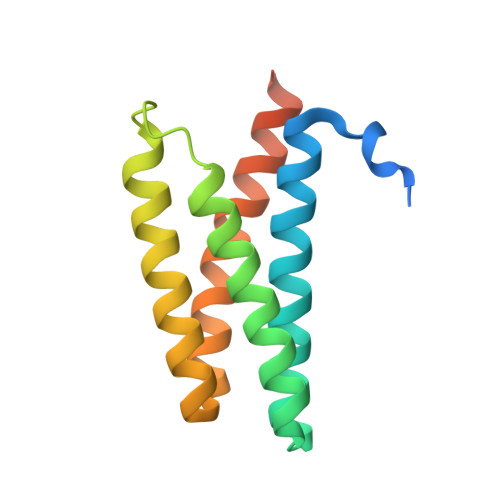
FliS/flagellin/FliW heterotrimer couples type III secretion and flagellin homeostasis.
Altegoer, F., Mukherjee, S., Steinchen, W., Bedrunka, P., Linne, U., Kearns, D.B., Bange, G.(2018) Sci Rep 8: 11552-11552
- PubMed: 30068950 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-29884-8
- Primary Citation Related Structures:
5MAW, 6GOW - PubMed Abstract:
Flagellin is amongst the most abundant proteins in flagellated bacterial species and constitutes the major building block of the flagellar filament. The proteins FliW and FliS serve in the post-transcriptional control of flagellin and guide the protein to the flagellar type III secretion system (fT3SS), respectively. Here, we present the high-resolution structure of FliS/flagellin heterodimer and show that FliS and FliW bind to opposing interfaces located at the N- and C-termini of flagellin. The FliS/flagellin/FliW heterotrimer is able to interact with FlhA-C suggesting that FliW and FliS are released during flagellin export. After release, FliW and FliS are recycled to execute a new round of post-transcriptional regulation and targeting. Taken together, our study provides a mechanism explaining how FliW and FliS synchronize the production of flagellin with the capacity of the fT3SS to secrete flagellin.
- LOEWE Center for Synthetic Microbiology & Dep. of Chemistry, Philipps University Marburg, Hans-Meerwein-Strasse 6, 35043, Marburg, Germany.
Organizational Affiliation: