Protein Crystallization via Sulfonatocalix[4]arene Dimethylammonium Complexation
Guagnini, F., Antonik, P.M., Rennie, M.L., Pinalli, R., Dalcanale, E., Crowley, P.B.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fucose-binding lectin protein | 90 | Ralstonia solanacearum | Mutation(s): 0 Gene Names: E7Z57_08365, RSP795_21825, RSP822_19650, RUN39_v1_50103 | 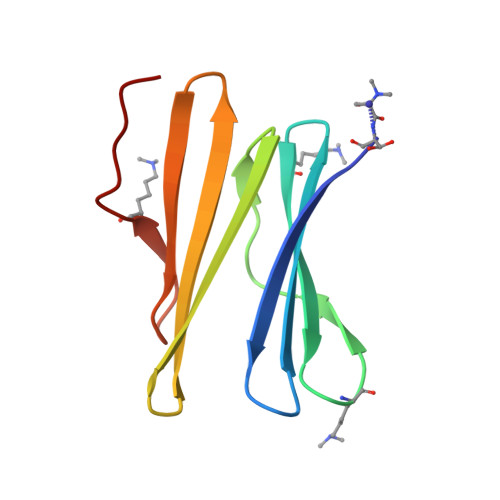 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A0S4TLR1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| T3Y (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | D [auth A] J [auth B] K [auth B] P [auth C] Q [auth C] | 25,26,27,28-tetrahydroxypentacyclo[19.3.1.1~3,7~.1~9,13~.1~15,19~]octacosa-1(25),3(28),4,6,9(27),10,12,15(26),16,18,21,23-dodecaene-5,11,17,23-tetrasulfonic acid C28 H24 O16 S4 JFYBCAFLVNKHHG-UHFFFAOYSA-N |  | ||
| PG4 Download:Ideal Coordinates CCD File | G [auth A] | TETRAETHYLENE GLYCOL C8 H18 O5 UWHCKJMYHZGTIT-UHFFFAOYSA-N |  | ||
| PEG Download:Ideal Coordinates CCD File | H [auth A], N [auth B], O [auth B], V [auth C] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| SO4 Download:Ideal Coordinates CCD File | U [auth C] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| GOL Download:Ideal Coordinates CCD File | E [auth A] F [auth A] L [auth B] M [auth B] S [auth C] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| EDO Download:Ideal Coordinates CCD File | I [auth A] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MLY Query on MLY | A, B, C | L-PEPTIDE LINKING | C8 H18 N2 O2 |  | LYS |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 53.92 | α = 90 |
| b = 67.57 | β = 90 |
| c = 87.75 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| Aimless | data scaling |
| PDB_EXTRACT | data extraction |
| MOSFLM | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Science Foundation Ireland | Ireland | 13/ERC/B2912 and 13/CDA/2168 |