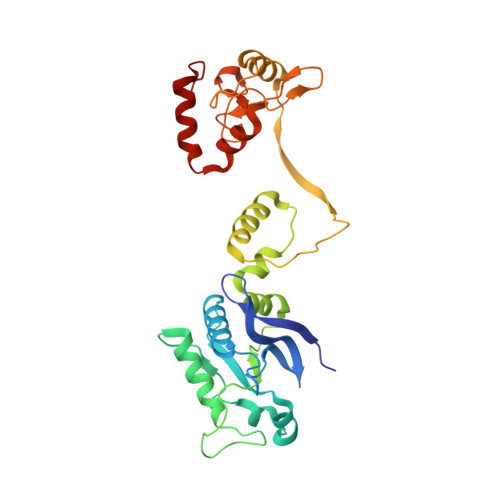
Activity and structure of EcoKMcrA.
Czapinska, H., Kowalska, M., Zagorskaite, E., Manakova, E., Slyvka, A., Xu, S.Y., Siksnys, V., Sasnauskas, G., Bochtler, M.(2018) Nucleic Acids Res 46: 9829-9841
- PubMed: 30107581 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gky731
- Primary Citation Related Structures:
6GHC - PubMed Abstract:
Escherichia coli McrA (EcoKMcrA) acts as a methylcytosine and hydroxymethylcytosine dependent restriction endonuclease. We present a biochemical characterization of EcoKMcrA that includes the first demonstration of its endonuclease activity, small angle X-ray scattering (SAXS) data, and a crystal structure of the enzyme in the absence of DNA. Our data indicate that EcoKMcrA dimerizes via the anticipated C-terminal HNH domains, which together form a single DNA binding site. The N-terminal domains are not homologous to SRA domains, do not interact with each other, and have separate DNA binding sites. Electrophoretic mobility shift assay (EMSA) and footprinting experiments suggest that the N-terminal domains can sense the presence and sequence context of modified cytosines. Pyrrolocytosine fluorescence data indicate no base flipping. In vitro, EcoKMcrA DNA endonuclease activity requires Mn2+ ions, is not strictly methyl dependent, and is not observed when active site variants of the enzyme are used. In cells, EcoKMcrA specifically restricts DNA that is modified in the correct sequence context. This activity is impaired by mutations of the nuclease active site, unless the enzyme is highly overexpressed.
- International Institute of Molecular and Cell Biology, Trojdena 4, 02-109 Warsaw, Poland.
Organizational Affiliation: