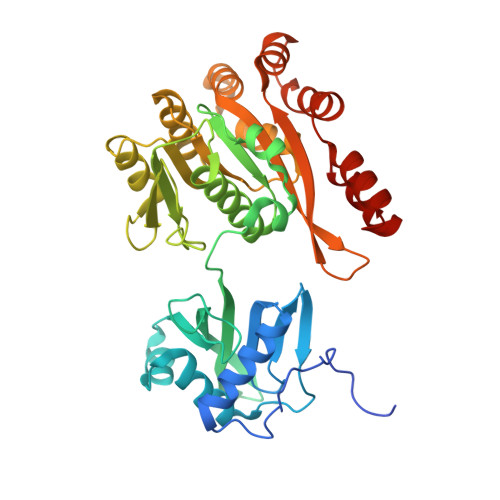
X-ray crystal structures of the type IVb secretion system DotB ATPases.
Prevost, M.S., Waksman, G.(2018) Protein Sci 27: 1464-1475
- PubMed: 29770512 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.3439
- Primary Citation Related Structures:
6GEB, 6GEF - PubMed Abstract:
Human infections by the intracellular bacterial pathogen Legionella pneumophila result in a severe form of pneumonia, the Legionnaire's disease. L. pneumophila utilizes a Type IVb secretion (T4bS) system termed "dot/icm" to secrete protein effectors to the host cytoplasm. The dot/icm system is powered at least in part by a functionally critical AAA+ ATPase, a protein called DotB, thought to belong to the VirB11 family of proteins. Here we present the crystal structure of DotB at 3.19 Å resolution, in its hexameric form. We observe that DotB is in fact a structural intermediate between VirB11 and PilT family proteins, with a PAS-like N-terminal domain coupled to a RecA-like C-terminal domain. It also shares critical structural elements only found in PilT. The structure also reveals two conformers, termed α and β, with an αβαβαβ configuration. The existence of α and β conformers in this class of proteins was confirmed by solving the structure of DotB from another bacterial pathogen, Yersinia, where, intriguingly, we observed an ααβααβ configuration. The two conformers co-exist regardless of the nucleotide-bound states of the proteins. Our investigation therefore reveals that these ATPases can adopt a wider range of conformational states than was known before, shedding new light on the extraordinary spectrum of conformations these ATPases can access to carry out their function. Overall, the structure of DotB provides a template for further rational drug design to develop more specific antibiotics to tackle Legionnaire's disease. PDB Code(s): Will; be; provided.
- Institute of Structural and Molecular Biology, University College London and Birkbeck, Malet Street, London, WC1E 7HX, United Kingdom.
Organizational Affiliation: