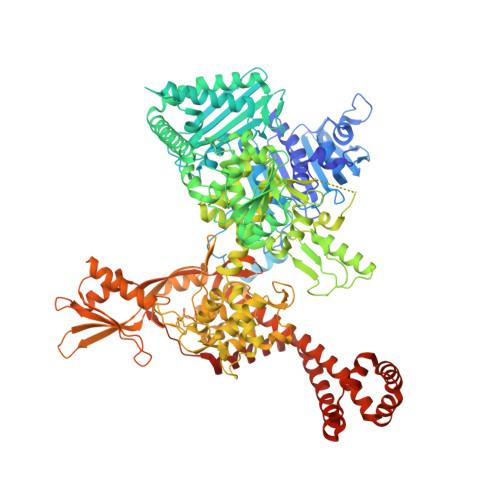
Overall Structures of Mycobacterium tuberculosis DNA Gyrase Reveal the Role of a Corynebacteriales GyrB-Specific Insert in ATPase Activity.
Petrella, S., Capton, E., Raynal, B., Giffard, C., Thureau, A., Bonnete, F., Alzari, P.M., Aubry, A., Mayer, C.(2019) Structure 27: 579-589.e5
- PubMed: 30744994 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2019.01.004
- Primary Citation Related Structures:
6GAU, 6GAV - PubMed Abstract:
Despite sharing common features, previous studies have shown that gyrases from different species have been modified throughout evolution to modulate their properties. Here, we report two crystal structures of Mycobacterium tuberculosis DNA gyrase, an apo and AMPPNP-bound form at 2.6-Å and 3.3-Å resolution, respectively. These structures provide high-resolution structural data on the quaternary organization and interdomain connections of a gyrase (full-length GyrB-GyrA57) 2 thus providing crucial inputs on this essential drug target. Together with small-angle X-ray scattering studies, they revealed an "extremely open" N-gate state, which persists even in the DNA-free gyrase-AMPPNP complex and an unexpected connection between the ATPase and cleavage core domains mediated by two Corynebacteriales-specific motifs, respectively the C-loop and DEEE-loop. We show that the C-loop participates in the stabilization of this open conformation, explaining why this gyrase has a lower ATPase activity. Our results image a conformational state which might be targeted for drug discovery.
- Unité de Microbiologie Structurale, Institut Pasteur, CNRS UMR 3528, 25 rue du Docteur Roux, 75724 Paris Cedex 15, France; Université Paris Diderot, Sorbonne Paris Cité, 75724 Paris Cedex 15, France. Electronic address: stephanie.petrella@pasteur.fr.
Organizational Affiliation: