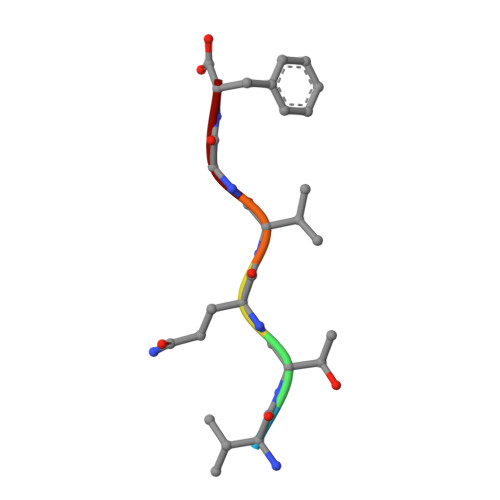
Structural Insights into Curli CsgA Cross-beta Fibril Architecture Inspire Repurposing of Anti-amyloid Compounds as Anti-biofilm Agents.
Perov, S., Lidor, O., Salinas, N., Golan, N., Tayeb-Fligelman, E., Deshmukh, M., Willbold, D., Landau, M.(2019) PLoS Pathog 15: e1007978-e1007978
- PubMed: 31469892 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1007978
- Primary Citation Related Structures:
6G8C, 6G8D, 6G8E, 6G9G - PubMed Abstract:
Curli amyloid fibrils secreted by Enterobacteriaceae mediate host cell adhesion and contribute to biofilm formation, thereby promoting bacterial resistance to environmental stressors. Here, we present crystal structures of amyloid-forming segments from the major curli subunit, CsgA, revealing steric zipper fibrils of tightly mated β-sheets, demonstrating a structural link between curli and human pathological amyloids. D-enantiomeric peptides, originally developed to interfere with Alzheimer's disease-associated amyloid-β, inhibited CsgA fibrillation and reduced biofilm formation in Salmonella typhimurium. Moreover, as previously shown, CsgA fibrils cross-seeded fibrillation of amyloid-β, providing support for the proposed structural resemblance and potential for cross-species amyloid interactions. The presented findings provide structural insights into amyloidogenic regions important for curli formation, suggest a novel strategy for disrupting amyloid-structured biofilms, and hypothesize on the formation of self-propagating prion-like species originating from a microbial source that could influence neurodegenerative diseases.
- Department of Biology, Technion-Israel Institute of Technology, Haifa, Israel.
Organizational Affiliation: