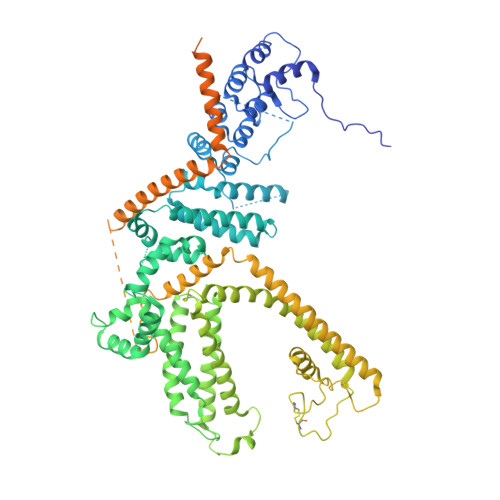
Electron cryo-microscopy structure of the canonical TRPC4 ion channel.
Vinayagam, D., Mager, T., Apelbaum, A., Bothe, A., Merino, F., Hofnagel, O., Gatsogiannis, C., Raunser, S.(2018) Elife 7
- PubMed: 29717981 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.36615
- Primary Citation Related Structures:
6G1K - PubMed Abstract:
Canonical transient receptor channels (TRPC) are non-selective cation channels. They are involved in receptor-operated Ca 2+ signaling and have been proposed to act as store-operated channels (SOC). Their malfunction is related to cardiomyopathies and their modulation by small molecules has been shown to be effective against renal cancer cells. The molecular mechanism underlying the complex activation and regulation is poorly understood. Here, we report the electron cryo-microscopy structure of zebrafish TRPC4 in its unliganded (apo), closed state at an overall resolution of 3.6 Å. The structure reveals the molecular architecture of the cation conducting pore, including the selectivity filter and lower gate. The cytoplasmic domain contains two key hubs that have been shown to interact with modulating proteins. Structural comparisons with other TRP channels give novel insights into the general architecture and domain organization of this superfamily of channels and help to understand their function and pharmacology.
- Department of Structural Biochemistry, Max Planck Institute of Molecular Physiology, Dortmund, Germany.
Organizational Affiliation: