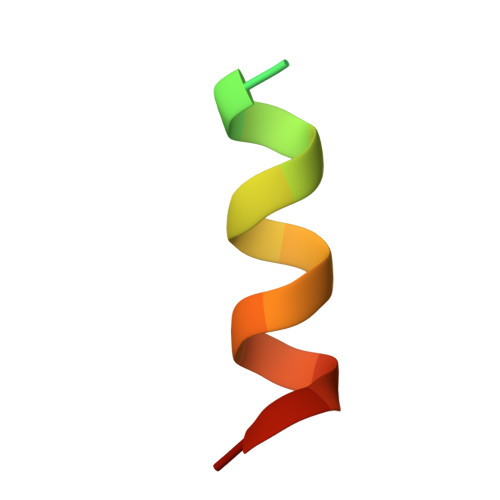
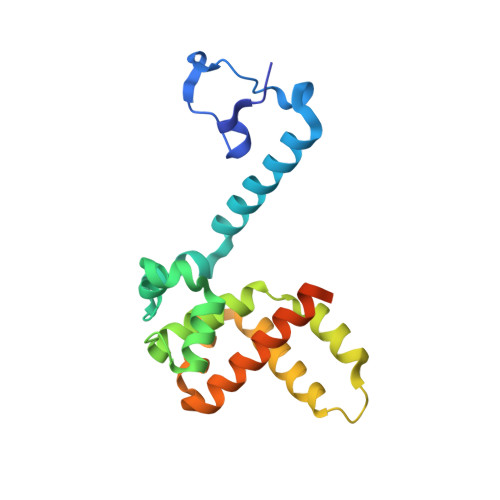
The Structure of the Human Respiratory Syncytial Virus M2-1 Protein Bound to the Interaction Domain of the Phosphoprotein P Defines the Orientation of the Complex.
Selvaraj, M., Yegambaram, K., Todd, E.J.A.A., Richard, C.A., Dods, R.L., Pangratiou, G.M., Trinh, C.H., Moul, S.L., Murphy, J.C., Mankouri, J., Eleouet, J.F., Barr, J.N., Edwards, T.A.(2018) mBio 9
- PubMed: 30425144 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mBio.01554-18
- Primary Citation Related Structures:
6G0Y - PubMed Abstract:
Human respiratory syncytial virus (HRSV) is a negative-stranded RNA virus that causes a globally prevalent respiratory infection, which can cause life-threatening illness, particularly in the young, elderly, and immunocompromised. HRSV multiplication depends on replication and transcription of the HRSV genes by the virus-encoded RNA-dependent RNA polymerase (RdRp). For replication, this complex comprises the phosphoprotein (P) and the large protein (L), whereas for transcription, the M2-1 protein is also required. M2-1 is recruited to the RdRp by interaction with P and also interacts with RNA at overlapping binding sites on the M2-1 surface, such that binding of these partners is mutually exclusive. The molecular basis for the transcriptional requirement of M2-1 is unclear, as is the consequence of competition between P and RNA for M2-1 binding, which is likely a critical step in the transcription mechanism. Here, we report the crystal structure at 2.4 Å of M2-1 bound to the P interaction domain, which comprises P residues 90 to 110. The P90-110 peptide is alpha helical, and its position on the surface of M2-1 defines the orientation of the three transcriptase components within the complex. The M2-1/P interface includes ionic, hydrophobic, and hydrogen bond interactions, and the critical contribution of these contacts to complex formation was assessed using a minigenome assay. The affinity of M2-1 for RNA and P ligands was quantified using fluorescence anisotropy, which showed high-affinity RNAs could outcompete P. This has important implications for the mechanism of transcription, particularly the events surrounding transcription termination and synthesis of poly(A) sequences. IMPORTANCE Human respiratory syncytial virus (HRSV) is a leading cause of respiratory illness, particularly in the young, elderly, and immunocompromised, and has also been linked to the development of asthma. HRSV replication depends on P and L, whereas transcription also requires M2-1. M2-1 interacts with P and RNA at overlapping binding sites; while these interactions are necessary for transcriptional activity, the mechanism of M2-1 action is unclear. To better understand HRSV transcription, we solved the crystal structure of M2-1 in complex with the minimal P interaction domain, revealing molecular details of the M2-1/P interface and defining the orientation of M2-1 within the tripartite complex. The M2-1/P interaction is relatively weak, suggesting high-affinity RNAs may displace M2-1 from the complex, providing the basis for a new model describing the role of M2-1 in transcription. Recently, the small molecules quercetin and cyclopamine have been used to validate M2-1 as a drug target.
- School of Molecular and Cellular Biology, University of Leeds, Leeds, United Kingdom.
Organizational Affiliation: