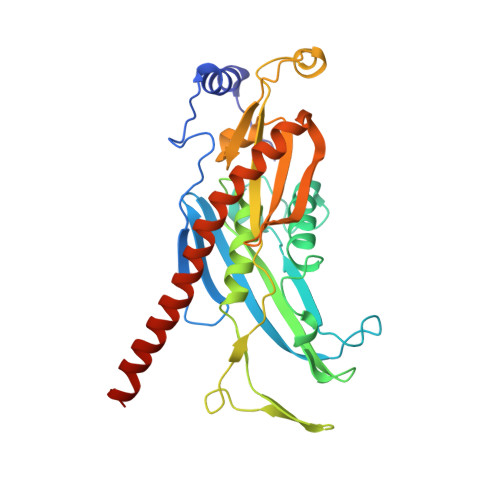
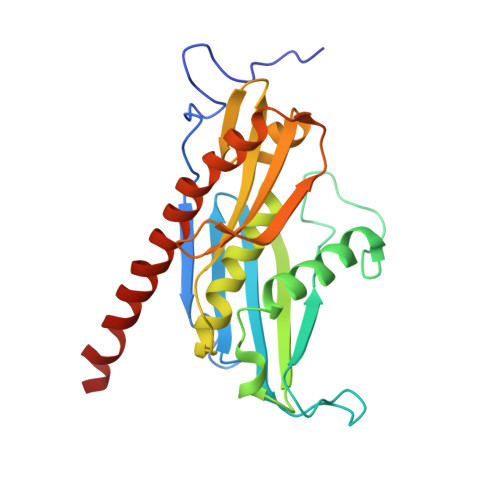
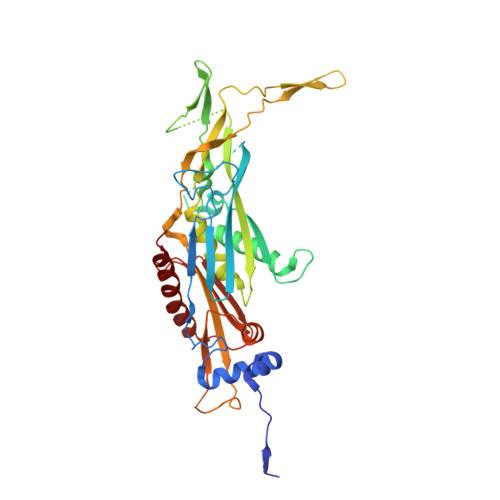
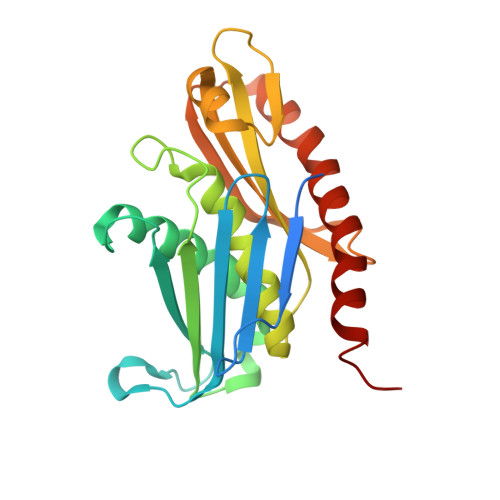
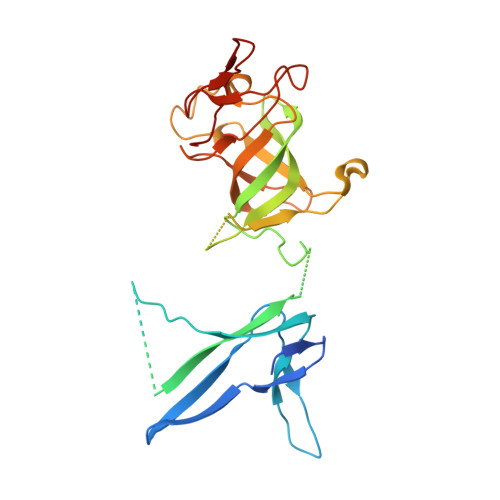
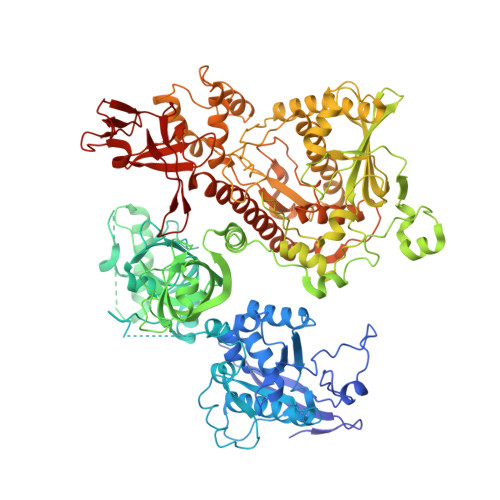
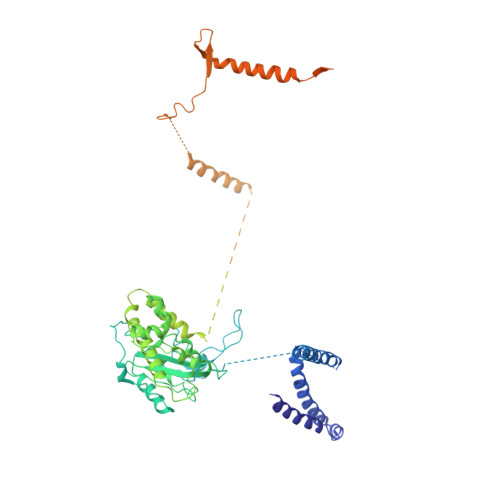
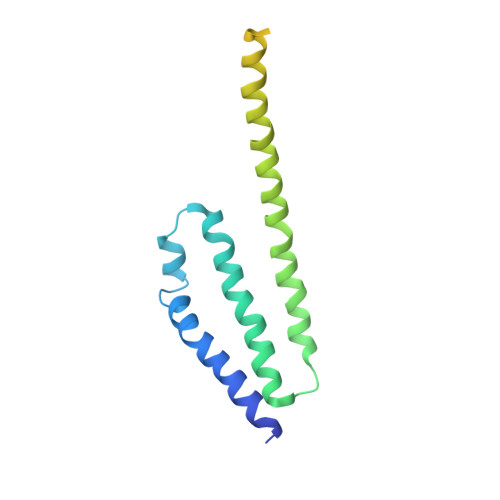
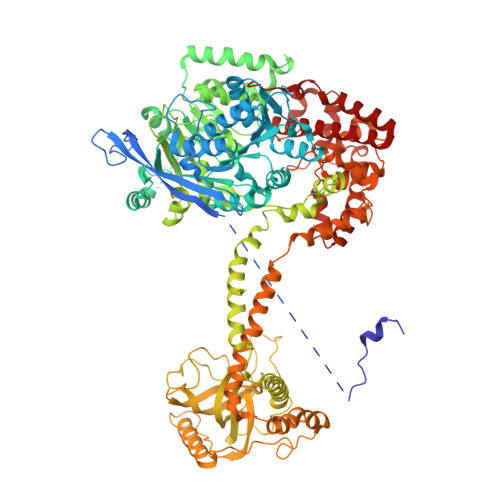
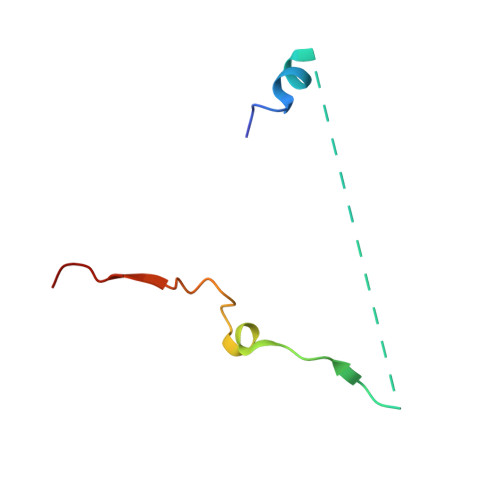
Structure of the nuclear exosome captured on a maturing preribosome.
Schuller, J.M., Falk, S., Fromm, L., Hurt, E., Conti, E.(2018) Science 360: 219-222
- PubMed: 29519915 Search on PubMed
- DOI: https://doi.org/10.1126/science.aar5428
- Primary Citation Related Structures:
6FSZ, 6FT6 - PubMed Abstract:
The RNA exosome complex processes and degrades a wide range of transcripts, including ribosomal RNAs (rRNAs). We used cryo-electron microscopy to visualize the yeast nuclear exosome holocomplex captured on a precursor large ribosomal subunit (pre-60 S ) during 7 S -to-5.8 S rRNA processing. The cofactors of the nuclear exosome are sandwiched between the ribonuclease core complex (Exo-10) and the remodeled "foot" structure of the pre-60 S particle, which harbors the 5.8 S rRNA precursor. The exosome-associated helicase Mtr4 recognizes the preribosomal substrate by docking to specific sites on the 25 S rRNA, captures the 3' extension of the 5.8 S rRNA, and channels it toward Exo-10. The structure elucidates how the exosome forms a structural and functional unit together with its massive pre-60 S substrate to process rRNA during ribosome maturation.
- Department of Structural Cell Biology, Max Planck Institute (MPI) for Biochemistry, Munich, Germany.
Organizational Affiliation: