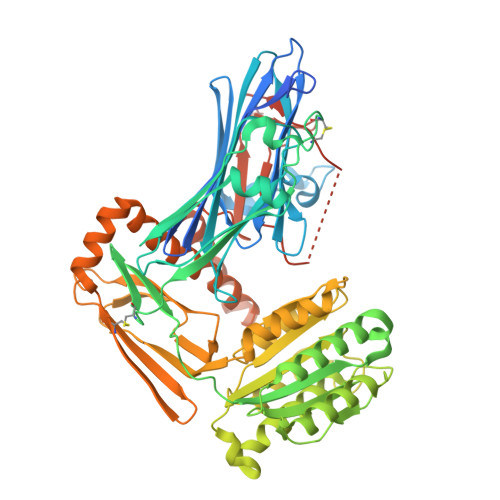
Inter-alpha-inhibitor heavy chain-1 has an integrin-like 3D structure mediating immune regulatory activities and matrix stabilization during ovulation
Briggs, D.C., Langford-Smith, A.W.W., Birchenough, H.L., Jowitt, T.A., Kielty, C.M., Enghild, J.J., Baldock, C., Milner, C.M., Day, A.J.(2020) J Biological Chem