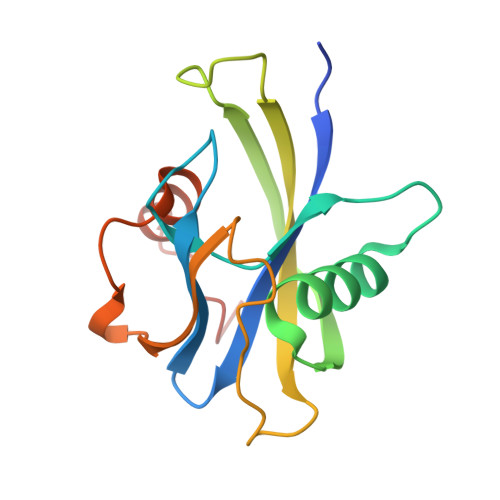
Crystal Structure and Substrate Specificity of the 8-oxo-dGTP Hydrolase NUDT1 from Arabidopsis thaliana.
Jemth, A.S., Scaletti, E., Carter, M., Helleday, T., Stenmark, P.(2019) Biochemistry 58: 887-899
- PubMed: 30614695 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.8b00950
- Primary Citation Related Structures:
6FL4 - PubMed Abstract:
Arabidopsis thaliana NUDT1 (AtNUDT1) belongs to the Nudix family of proteins, which have a diverse range of substrates, including oxidized nucleotides such as 8-oxo-dGTP. The hydrolysis of oxidized dNTPs is highly important as it prevents their incorporation into DNA, thus preventing mutations and DNA damage. AtNUDT1 is the sole Nudix enzyme from A. thaliana shown to have activity against 8-oxo-dGTP. We present the structure of AtNUDT1 in complex with 8-oxo-dGTP. Structural comparison with bacterial and human homologues reveals a conserved overall fold. Analysis of the 8-oxo-dGTP binding mode shows that the residues Asn76 and Ser89 interact with the O8 atom of the substrate, a feature not observed in structures of protein homologues solved to date. Kinetic analysis of wild-type and mutant AtNUDT1 confirmed that these active site residues influence 8-oxo-dGTP hydrolysis. A recent study showed that AtNUDT1 is also able to hydrolyze terpene compounds. The diversity of reactions catalyzed by AtNUDT1 suggests that this Nudix enzyme from higher plants has evolved in a manner distinct to those from other organisms.
- Science for Life Laboratory, Division of Translational Medicine and Chemical Biology, Department of Medical Biochemistry and Biophysics , Karolinska Institutet , Stockholm S-171 21 , Sweden.
Organizational Affiliation: