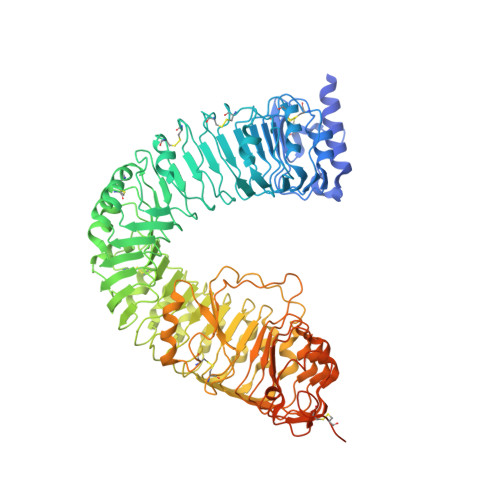
Mechanistic basis for the activation of plant membrane receptor kinases by SERK-family coreceptors.
Hohmann, U., Santiago, J., Nicolet, J., Olsson, V., Spiga, F.M., Hothorn, L.A., Butenko, M.A., Hothorn, M.(2018) Proc Natl Acad Sci U S A 115: 3488-3493
- PubMed: 29531026 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1714972115
- Primary Citation Related Structures:
6FIF - PubMed Abstract:
Plant-unique membrane receptor kinases with leucine-rich repeat ectodomains (LRR-RKs) can sense small molecule, peptide, and protein ligands. Many LRR-RKs require SERK-family coreceptor kinases for high-affinity ligand binding and receptor activation. How one coreceptor can contribute to the specific binding of distinct ligands and activation of different LRR-RKs is poorly understood. Here we quantitatively analyze the contribution of SERK3 to ligand binding and activation of the brassinosteroid receptor BRI1 and the peptide hormone receptor HAESA. We show that while the isolated receptors sense their respective ligands with drastically different binding affinities, the SERK3 ectodomain binds the ligand-associated receptors with very similar binding kinetics. We identify residues in the SERK3 N-terminal capping domain, which allow for selective steroid and peptide hormone recognition. In contrast, residues in the SERK3 LRR core form a second, constitutive receptor-coreceptor interface. Genetic analyses of protein chimera between BRI1 and SERK3 define that signaling-competent complexes are formed by receptor-coreceptor heteromerization in planta. A functional BRI1-HAESA chimera suggests that the receptor activation mechanism is conserved among different LRR-RKs, and that their signaling specificity is encoded in the kinase domain of the receptor. Our work pinpoints the relative contributions of receptor, ligand, and coreceptor to the formation and activation of SERK-dependent LRR-RK signaling complexes regulating plant growth and development.
- Structural Plant Biology Laboratory, Department of Botany and Plant Biology, University of Geneva, 1211 Geneva, Switzerland.
Organizational Affiliation: