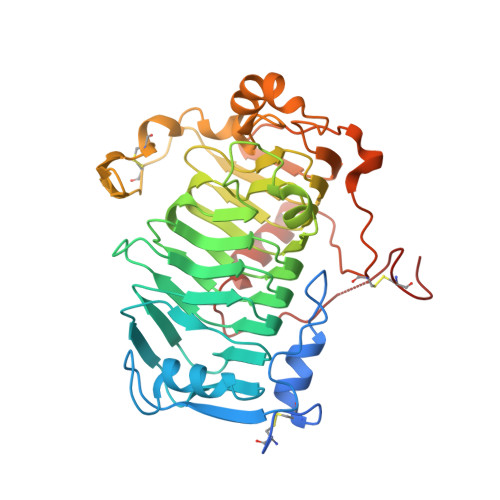
Periplasmic depolymerase provides insight into ABC transporter-dependent secretion of bacterial capsular polysaccharides.
Liston, S.D., McMahon, S.A., Le Bas, A., Suits, M.D.L., Naismith, J.H., Whitfield, C.(2018) Proc Natl Acad Sci U S A 115: E4870-E4879
- PubMed: 29735649 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1801336115
- Primary Citation Related Structures:
6FI2 - PubMed Abstract:
Capsules are surface layers of hydrated capsular polysaccharides (CPSs) produced by many bacteria. The human pathogen Salmonella enterica serovar Typhi produces "Vi antigen" CPS, which contributes to virulence. In a conserved strategy used by bacteria with diverse CPS structures, translocation of Vi antigen to the cell surface is driven by an ATP-binding cassette (ABC) transporter. These transporters are engaged in heterooligomeric complexes proposed to form an enclosed translocation conduit to the cell surface, allowing the transporter to power the entire process. We identified Vi antigen biosynthesis genetic loci in genera of the Burkholderiales , which are paradoxically distinguished from S. Typhi by encoding VexL, a predicted pectate lyase homolog. Biochemical analyses demonstrated that VexL is an unusual metal-independent endolyase with an acidic pH optimum that is specific for O-acetylated Vi antigen. A 1.22-Å crystal structure of the VexL-Vi antigen complex revealed features which distinguish common secreted catabolic pectate lyases from periplasmic VexL, which participates in cell-surface assembly. VexL possesses a right-handed parallel β-superhelix, of which one face forms an electropositive glycan-binding groove with an extensive hydrogen bonding network that includes Vi antigen acetyl groups and confers substrate specificity. VexL provided a probe to interrogate conserved features of the ABC transporter-dependent export model. When introduced into S Typhi, VexL localized to the periplasm and degraded Vi antigen. In contrast, a cytosolic derivative had no effect unless export was disrupted. These data provide evidence that CPS assembled in ABC transporter-dependent systems is actually exposed to the periplasm during envelope translocation.
- Department of Molecular and Cellular Biology, University of Guelph, Guelph, ON N1G 2W1, Canada.
Organizational Affiliation: