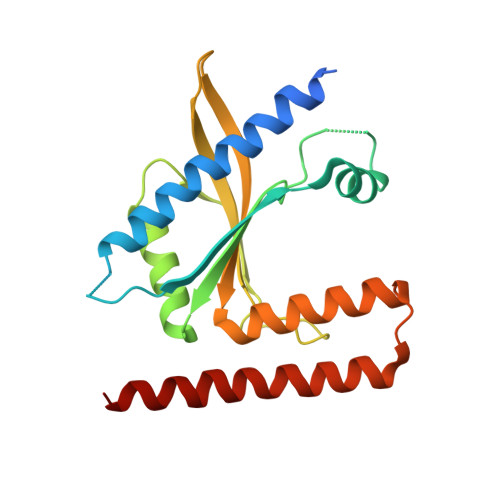
Structural and mechanistic divergence of the small (p)ppGpp synthetases RelP and RelQ.
Steinchen, W., Vogt, M.S., Altegoer, F., Giammarinaro, P.I., Horvatek, P., Wolz, C., Bange, G.(2018) Sci Rep 8: 2195-2195
- PubMed: 29391580 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-20634-4
- Primary Citation Related Structures:
6FGJ, 6FGK, 6FGX - PubMed Abstract:
The nutritional alarmones ppGpp and pppGpp (collectively: (p)ppGpp) are nucleotide-based second messengers enabling bacteria to respond to environmental and stress conditions. Several bacterial species contain two highly homologous (p)ppGpp synthetases named RelP (SAS2, YwaC) and RelQ (SAS1, YjbM). It is established that RelQ forms homotetramers that are subject to positive allosteric regulation by pppGpp, but structural and mechanistic insights into RelP lack behind. Here we present a structural and mechanistic characterization of RelP. In stark contrast to RelQ, RelP is not allosterically regulated by pppGpp and displays a different enzyme kinetic behavior. This discrepancy is evoked by different conformational properties of the guanosine-substrate binding site (G-Loop) of both proteins. Our study shows how minor structural divergences between close homologues result in new functional features during the course of molecular evolution.
- Philipps-University Marburg, LOEWE Center for Synthetic Microbiology & Department of Chemistry, Hans-Meerwein-Straße, 35043 Marburg, Germany. wieland.steinchen@synmikro.uni-marburg.de.
Organizational Affiliation: