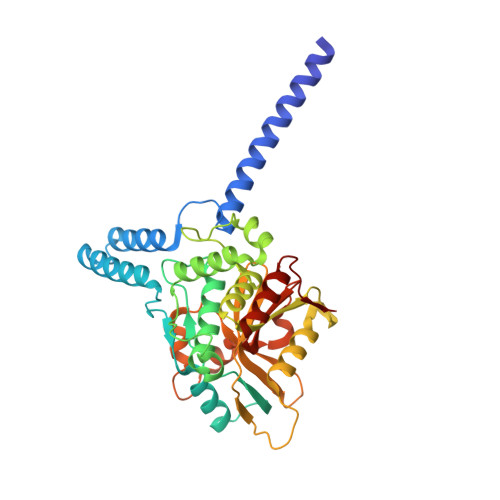
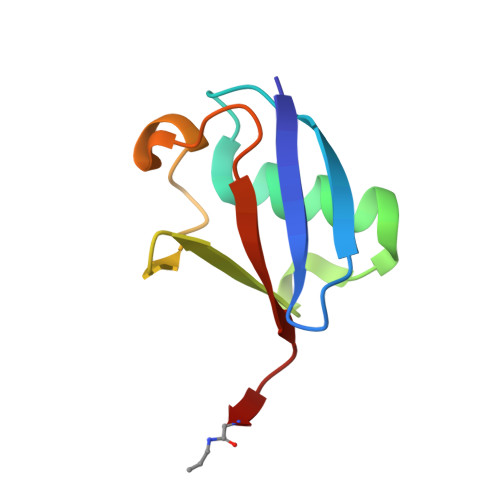
Discovery and Characterization of ZUFSP/ZUP1, a Distinct Deubiquitinase Class Important for Genome Stability.
Kwasna, D., Abdul Rehman, S.A., Natarajan, J., Matthews, S., Madden, R., De Cesare, V., Weidlich, S., Virdee, S., Ahel, I., Gibbs-Seymour, I., Kulathu, Y.(2018) Mol Cell 70: 150-164.e6
- PubMed: 29576527 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2018.02.023
- Primary Citation Related Structures:
6FGE - PubMed Abstract:
Deubiquitinating enzymes (DUBs) are important regulators of ubiquitin signaling. Here, we report the discovery of deubiquitinating activity in ZUFSP/C6orf113. High-resolution crystal structures of ZUFSP in complex with ubiquitin reveal several distinctive features of ubiquitin recognition and catalysis. Our analyses reveal that ZUFSP is a novel DUB with no homology to any known DUBs, leading us to classify ZUFSP as the seventh DUB family. Intriguingly, the minimal catalytic domain does not cleave polyubiquitin. We identify two ubiquitin binding domains in ZUFSP: a ZHA (ZUFSP helical arm) that binds to the distal ubiquitin and an atypical UBZ domain in ZUFSP that binds to polyubiquitin. Importantly, both domains are essential for ZUFSP to selectively cleave K63-linked polyubiquitin. We show that ZUFSP localizes to DNA lesions, where it plays an important role in genome stability pathways, functioning to prevent spontaneous DNA damage and also promote cellular survival in response to exogenous DNA damage.
- MRC Protein Phosphorylation & Ubiquitylation Unit, School of Life Sciences, University of Dundee, Dow Street, Dundee DD1 5EH, UK.
Organizational Affiliation: