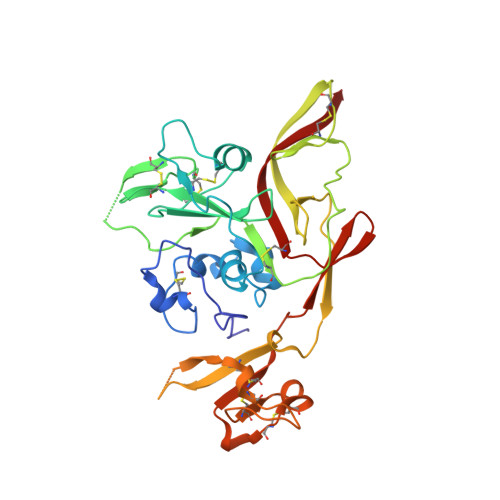
Shielding and activation of a viral membrane fusion protein.
Halldorsson, S., Li, S., Li, M., Harlos, K., Bowden, T.A., Huiskonen, J.T.(2018) Nat Commun 9: 349-349
- PubMed: 29367607 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02789-2
- Primary Citation Related Structures:
6F8P, 6F9B, 6F9C, 6F9D, 6F9E, 6F9F - PubMed Abstract:
Entry of enveloped viruses relies on insertion of hydrophobic residues of the viral fusion protein into the host cell membrane. However, the intermediate conformations during fusion remain unknown. Here, we address the fusion mechanism of Rift Valley fever virus. We determine the crystal structure of the Gn glycoprotein and fit it with the Gc fusion protein into cryo-electron microscopy reconstructions of the virion. Our analysis reveals how the Gn shields the hydrophobic fusion loops of the Gc, preventing premature fusion. Electron cryotomography of virions interacting with membranes under acidic conditions reveals how the fusogenic Gc is activated upon removal of the Gn shield. Repositioning of the Gn allows extension of Gc and insertion of fusion loops in the outer leaflet of the target membrane. These data show early structural transitions that enveloped viruses undergo during host cell entry and indicate that analogous shielding mechanisms are utilized across diverse virus families.
- Division of Structural Biology, Wellcome Centre for Human Genetics, University of Oxford, Roosevelt Drive, Oxford, OX3 7BN, UK.
Organizational Affiliation: