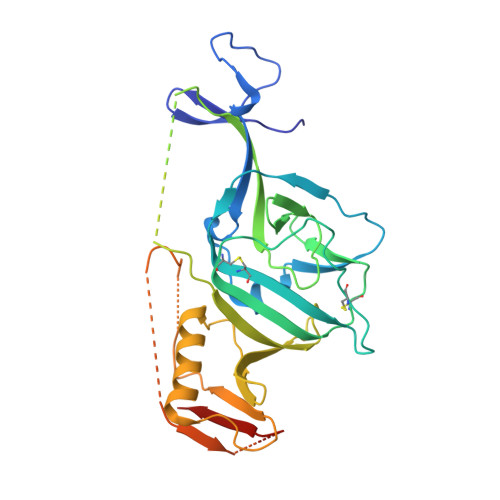
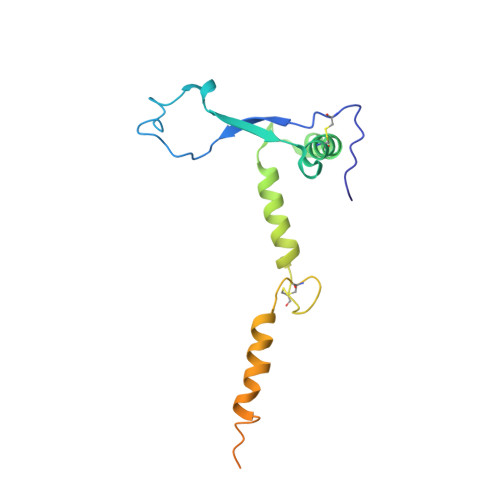
Target Identification and Mode of Action of Four Chemically Divergent Drugs against Ebolavirus Infection.
Ren, J., Zhao, Y., Fry, E.E., Stuart, D.I.(2018) J Med Chem 61: 724-733
- PubMed: 29272110
- DOI: https://doi.org/10.1021/acs.jmedchem.7b01249
- Primary Citation Related Structures:
6F5U, 6F6I, 6F6N, 6F6S - PubMed Abstract:
Here, we show that four chemically divergent approved drugs reported to inhibit Ebolavirus infection, benztropine, bepridil, paroxetine and sertraline, directly interact with the Ebolavirus glycoprotein. Binding of these drugs destabilizes the protein, suggesting that this may be the mechanism of inhibition, as reported for the anticancer drug toremifene and the painkiller ibuprofen, which bind in the same large cavity on the glycoprotein. Crystal structures show that the position of binding and the mode of interaction within the pocket vary significantly between these compounds. The binding constants (K d ) determined by thermal shift assay correlate with the protein-inhibitor interactions as well as with the antiviral activities determined by virus cell entry assays, supporting the hypothesis that these drugs inhibit viral entry by binding the glycoprotein and destabilizing the prefusion conformation. Details of the protein-inhibitor interactions of these complexes and their relation with binding affinity may facilitate the design of more potent inhibitors.
- Division of Structural Biology, University of Oxford , The Henry Wellcome Building for Genomic Medicine, Headington, Oxford, OX3 7BN, U.K.
Organizational Affiliation: