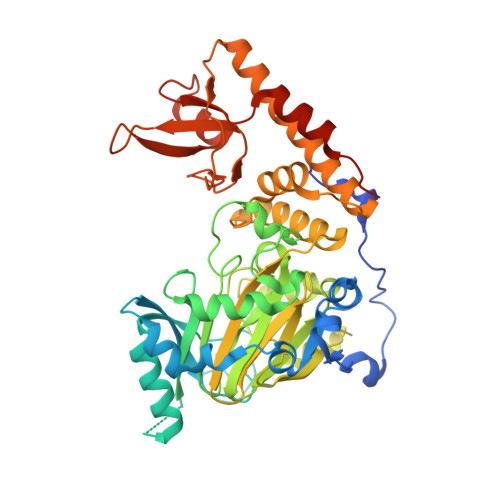
Structural Basis of Histone Demethylase KDM6B Histone 3 Lysine 27 Specificity.
Jones, S.E., Olsen, L., Gajhede, M.(2018) Biochemistry 57: 585-592
- PubMed: 29220567 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.7b01152
- Primary Citation Related Structures:
5OY3, 6F6D - PubMed Abstract:
KDM subfamily 6 enzymes KDM6A and KDM6B specifically catalyze demethylation of di- and trimethylated lysine on histone 3 lysine 27 (H3K27me3/2) and play an important role in repression of developmental genes. Despite identical amino acid sequence in the immediate surroundings of H3K9me3/2 (ARKS), the enzymes do not catalyze demethylation of this general marker of repression. To address this question for KDM6B, we used computational methods to identify H3(17-33)-derived peptides with improved binding affinity that would allow co-crystallization with the catalytic core of human KDM6B (ccKDM6B). A total of five peptides were identified, and their IC 50 values were determined in a matrix-assisted laser desorption ionization time-of-flight-based assay. Despite none of the peptides showing affinity significantly higher than that of the H3(17-33) peptide, it was possible to co-crystallize ccKDM6B with a H3(17-33)A21M peptide. This structure reveals the interactions between the KDM6B zinc binding domain and the H3(17-23) region. Although KDM6A and KDM6B differ in primary sequence, particularly in the H3L20 binding pocket of the zinc binding domains, where two histidines in KDM6A have been replaced by a glutamate and a tyrosine, they bind H3(17-23) in a very similar fashion. This structure shows that KDM6B, in analogy with KDM6A, also uses the zinc binding domain to achieve H3K27me3/me2 specificity. The histidine to glutamine substitution at amino acid position 1564 in the KDM6B zinc binding domain can further explain why KDM6B binds substrates with an affinity higher than that of KDM6A.
- Biostructural Research, Department of Drug Design and Pharmacology, Faculty of Health and Medical Sciences, University of Copenhagen , Jagtvej 162, 2100 Copenhagen, Denmark.
Organizational Affiliation: