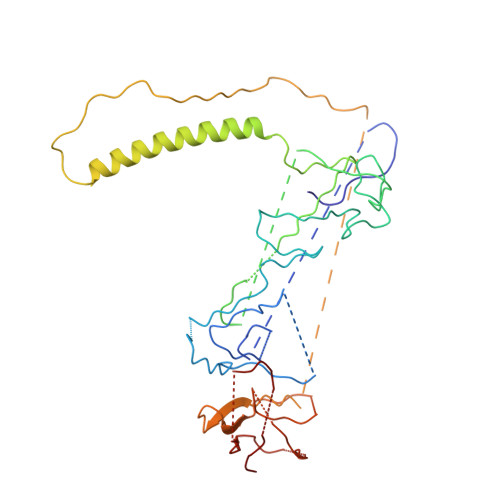
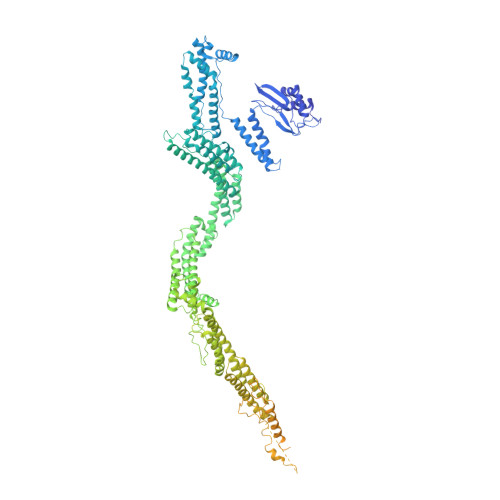
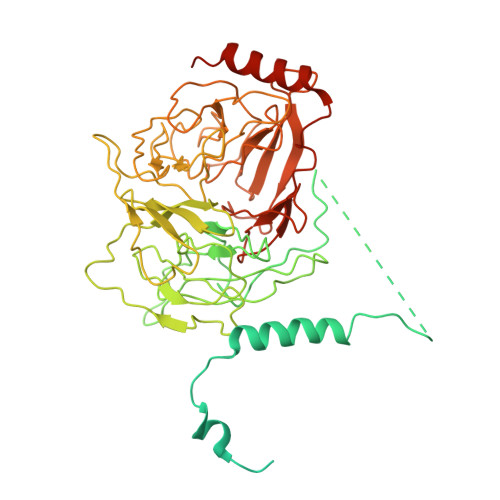
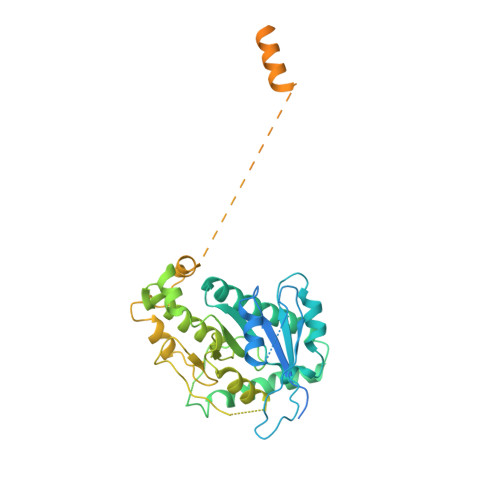
Cryo-EM shows how dynactin recruits two dyneins for faster movement.
Urnavicius, L., Lau, C.K., Elshenawy, M.M., Morales-Rios, E., Motz, C., Yildiz, A., Carter, A.P.(2018) Nature 554: 202-206
- PubMed: 29420470 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature25462
- Primary Citation Related Structures:
5OWO, 6F1T, 6F1U, 6F1V, 6F1Y, 6F1Z, 6F38, 6F3A - PubMed Abstract:
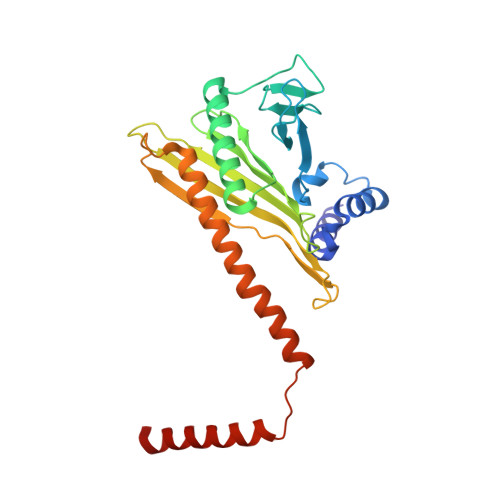
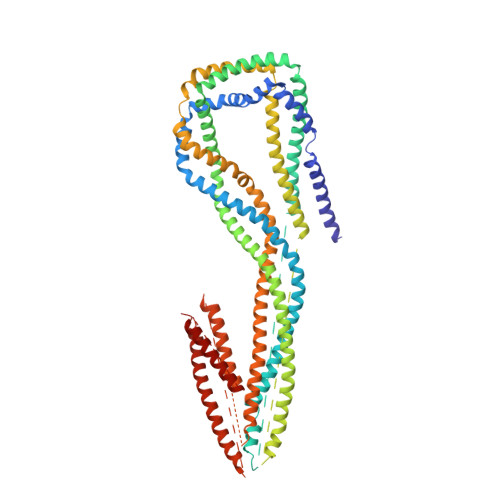
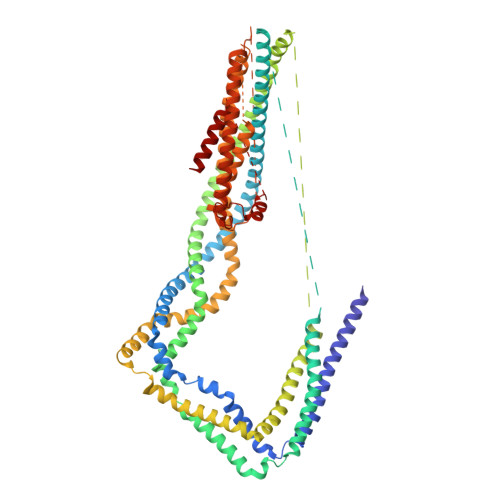
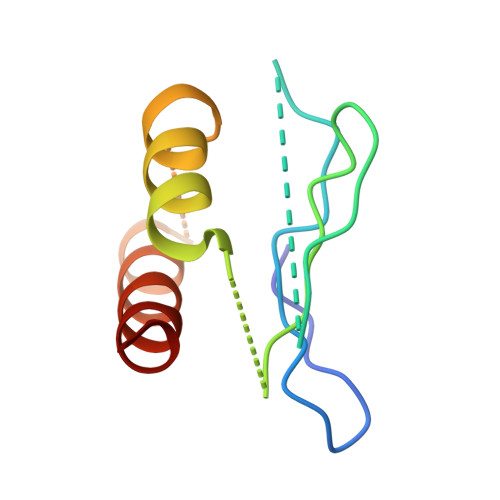
Dynein and its cofactor dynactin form a highly processive microtubule motor in the presence of an activating adaptor, such as BICD2. Different adaptors link dynein and dynactin to distinct cargoes. Here we use electron microscopy and single-molecule studies to show that adaptors can recruit a second dynein to dynactin. Whereas BICD2 is biased towards recruiting a single dynein, the adaptors BICDR1 and HOOK3 predominantly recruit two dyneins. We find that the shift towards a double dynein complex increases both the force and speed of the microtubule motor. Our 3.5 Å resolution cryo-electron microscopy reconstruction of a dynein tail-dynactin-BICDR1 complex reveals how dynactin can act as a scaffold to coordinate two dyneins side-by-side. Our work provides a structural basis for understanding how diverse adaptors recruit different numbers of dyneins and regulate the motile properties of the dynein-dynactin transport machine.
- Medical Research Council Laboratory of Molecular Biology, Division of Structural Studies, Francis Crick Avenue, Cambridge CB2 0QH, UK.
Organizational Affiliation: