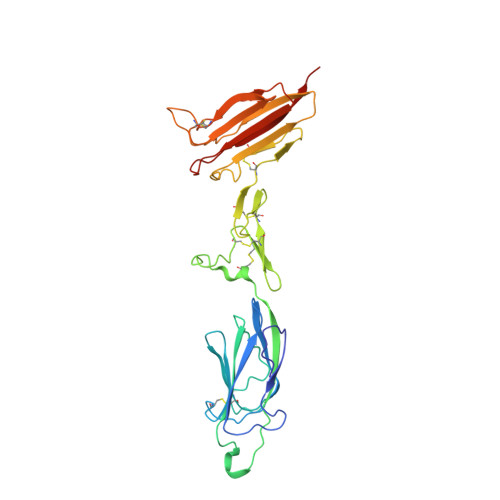
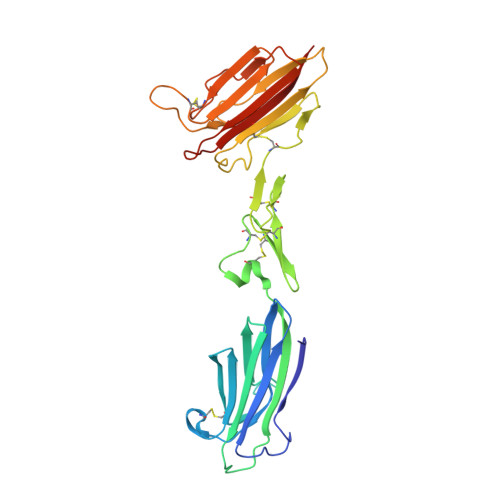
Structure of the C1r-C1s interaction of the C1 complex of complement activation.
Almitairi, J.O.M., Venkatraman Girija, U., Furze, C.M., Simpson-Gray, X., Badakshi, F., Marshall, J.E., Schwaeble, W.J., Mitchell, D.A., Moody, P.C.E., Wallis, R.(2018) Proc Natl Acad Sci U S A 115: 768-773
- PubMed: 29311313 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1718709115
- Primary Citation Related Structures:
6F1C, 6F1D, 6F1H, 6F39 - PubMed Abstract:
The multiprotein complex C1 initiates the classical pathway of complement activation on binding to antibody-antigen complexes, pathogen surfaces, apoptotic cells, and polyanionic structures. It is formed from the recognition subcomponent C1q and a tetramer of proteases C1r 2 C1s 2 as a Ca 2+ -dependent complex. Here we have determined the structure of a complex between the CUB1-EGF-CUB2 fragments of C1r and C1s to reveal the C1r-C1s interaction that forms the core of C1. Both fragments are L-shaped and interlock to form a compact antiparallel heterodimer with a Ca 2+ from each subcomponent at the interface. Contacts, involving all three domains of each protease, are more extensive than those of C1r or C1s homodimers, explaining why heterocomplexes form preferentially. The available structural and biophysical data support a model of C1r 2 C1s 2 in which two C1r-C1s dimers are linked via the catalytic domains of C1r. They are incompatible with a recent model in which the N-terminal domains of C1r and C1s form a fixed tetramer. On binding to C1q, the proteases become more compact, with the C1r-C1s dimers at the center and the six collagenous stems of C1q arranged around the perimeter. Activation is likely driven by separation of the C1r-C1s dimer pairs when C1q binds to a surface. Considerable flexibility in C1s likely facilitates C1 complex formation, activation of C1s by C1r, and binding and activation of downstream substrates C4 and C4b-bound C2 to initiate the reaction cascade.
- Department of Infection, Immunity, and Inflammation, University of Leicester, Leicester LE1 9HN, United Kingdom.
Organizational Affiliation: