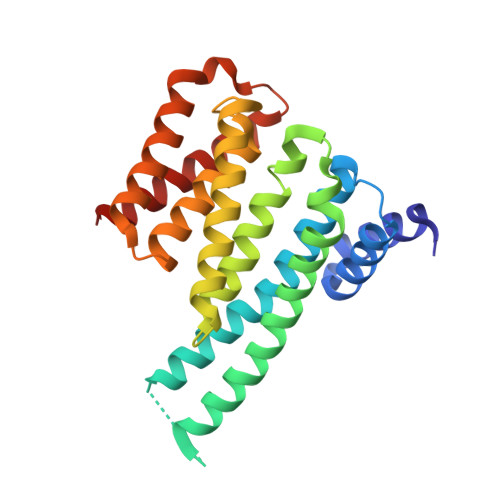
Structural characterization of 14-3-3 zeta in complex with the human Son of sevenless homolog 1 (SOS1).
Ballone, A., Centorrino, F., Wolter, M., Ottmann, C.(2018) J Struct Biol 202: 210-215
- PubMed: 29408703
- DOI: https://doi.org/10.1016/j.jsb.2018.01.011
- Primary Citation of Related Structures:
6F08 - PubMed Abstract:
The deviant Ras activation machinery is found in approximately 30% of all human cancers. SOS1 is an important protagonist of this pathway that plays a key-role in aberrant cell proliferation and differentiation. Interaction of SOS1 with 14-3-3 proteins modulates SOS1 activity in Ras-MAPK signaling. In the present study, we analyze the 14-3-3/SOS1 protein-protein interaction (PPI) by different biochemical assays and report the high resolution crystal structure of a 13-mer motif of SOS1 bound to 14-3-3ζ. These structural and functional insights are important for the evaluation of this PPI interface for small-molecule stabilization as a new starting point for modulating the Ras-Raf-MAPK pathway.
- Laboratory of Chemical Biology, Department of Biomedical Engineering and Institute for Molecular Systems, Eindhoven University of Technology, Eindhoven 5600 MB, The Netherlands.
Organizational Affiliation: