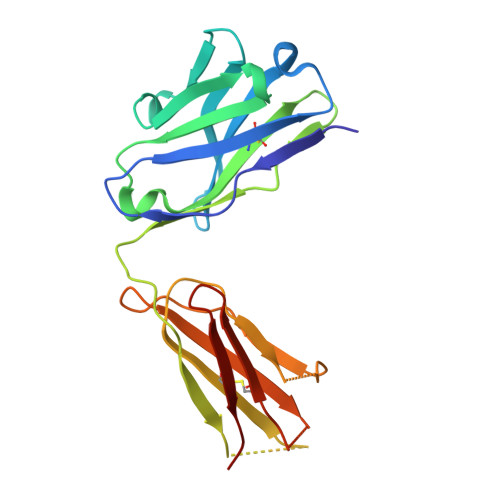
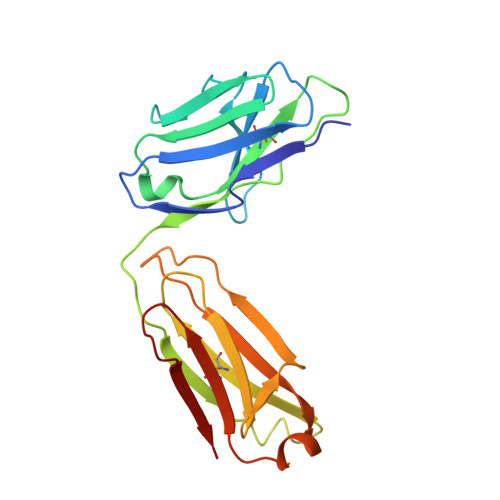
Human Norovirus Neutralized by a Monoclonal Antibody Targeting the Histo-Blood Group Antigen Pocket.
Koromyslova, A.D., Morozov, V.A., Hefele, L., Hansman, G.S.(2019) J Virol 93
- PubMed: 30541855
- DOI: https://doi.org/10.1128/JVI.02174-18
- Primary Citation of Related Structures:
6EWB - PubMed Abstract:
Temporal changes in the GII.4 human norovirus capsid sequences occasionally result in the emergence of genetic variants capable of causing new epidemics. The persistence of GII.4 is believed to be associated with the recognition of numerous histo-blood group antigen (HBGA) types and antigenic drift. We found that one of the earliest known GII.4 isolates (in 1974) and a more recent epidemic GII.4 variant (in 2012) had varied norovirus-specific monoclonal antibody (MAb) reactivities but similar HBGA binding profiles. To better understand the binding interaction of one MAb (10E9) that had varied reactivity with these GII.4 variants, we determined the X-ray crystal structure of the NSW-2012 GII.4 P domain 10E9 Fab complex. We showed that the 10E9 Fab interacted with conserved and variable residues, which could be associated with antigenic drift. Interestingly, the 10E9 Fab binding pocket partially overlapped the HBGA pocket and had direct competition for conserved HBGA binding residues (i.e., Arg345 and Tyr444). Indeed, the 10E9 MAb blocked norovirus virus-like particles (VLPs) from binding to several sources of HBGAs. Moreover, the 10E9 antibody completely abolished virus replication in the human norovirus intestinal enteroid cell culture system. Our new findings provide the first direct evidence that competition for GII.4 HBGA binding residues and steric obstruction could lead to norovirus neutralization. On the other hand, the 10E9 MAb recognized residues flanking the HBGA pocket, which are often substituted as the virus evolves. This mechanism of antigenic drift likely influences herd immunity and impedes the possibility of acquiring broadly reactive HBGA-blocking antibodies. IMPORTANCE The emergence of new epidemic GII.4 norovirus variants is thought to be associated with changes in antigenicity and HBGA binding capacity. Here, we show that HBGA binding profiles remain unchanged between the 1974 and 2012 GII.4 variants, whereas these variants showed various levels of reactivity against a panel of GII.4 MAbs. We identified a MAb that bound at the HBGA pocket, blocked norovirus VLPs from binding to HBGAs, and neutralized norovirus virions in the cell culture system. Raised against a GII.4 2006 strain, this MAb was unreactive to a GII.4 1974 isolate but was able to neutralize the newer 2012 strain, which has important implications for vaccine design. Altogether, these new findings suggest that the amino acid variations surrounding the HBGA pocket lead to temporal changes in antigenicity without affecting the ability of GII.4 variants to bind HBGAs, which are known cofactors for infection.
- Schaller Research Group at the University of Heidelberg and the DKFZ, Heidelberg, Germany a.koromyslova@dkfz.de g.hansman@dkfz.de.
Organizational Affiliation: