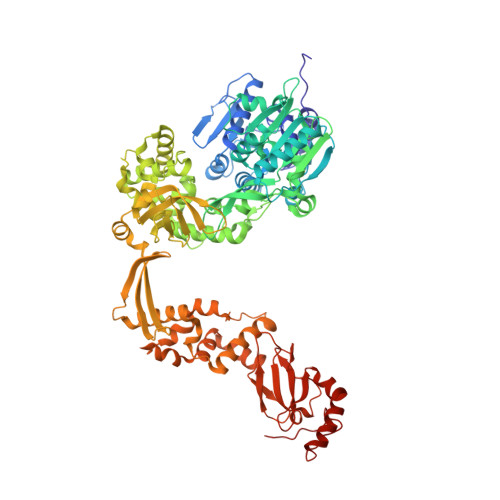
Crystal Structure of the Escherichia coli DExH-Box NTPase HrpB.
Pietrzyk-Brzezinska, A.J., Absmeier, E., Klauck, E., Wen, Y., Antelmann, H., Wahl, M.C.(2018) Structure 26: 1462-1473.e4
- PubMed: 30174149 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2018.07.013
- Primary Citation Related Structures:
6EUD - PubMed Abstract:
Eukaryotic DExH-box proteins are important post-transcriptional gene regulators, many of which employ RNA-stimulated nucleoside triphosphatase activity to remodel RNAs or ribonucleoprotein complexes. However, bacterial DExH-box proteins are structurally and functionally poorly characterized. We report the crystal structure of the Escherichia coli DExH-box protein HrpB. A globular head is composed of dual RecA, winged-helix, helical bundle and oligonucleotide/oligosaccharide-binding domains, resembling a compact version of eukaryotic DExH-box proteins. Additionally, HrpB harbors a C-terminal region not found in proteins with known structure, which bestows the protein with unique interaction potential. Interaction and activity assays showed that the protein binds RNA but not DNA, hydrolyzes all nucleoside triphosphates in an RNA-stimulated manner, but does not unwind diverse model RNAs in vitro. These observations can be rationalized by detailed comparisons with structurally characterized eukaryotic DExH-box proteins. Comparative phenotypic analyses of an E. coli hrpB knockout mutant suggested diverse functions of HrpB homologs in different bacteria.
- Freie Universität Berlin, Laboratory of Structural Biochemistry, 14195 Berlin, Germany.
Organizational Affiliation: