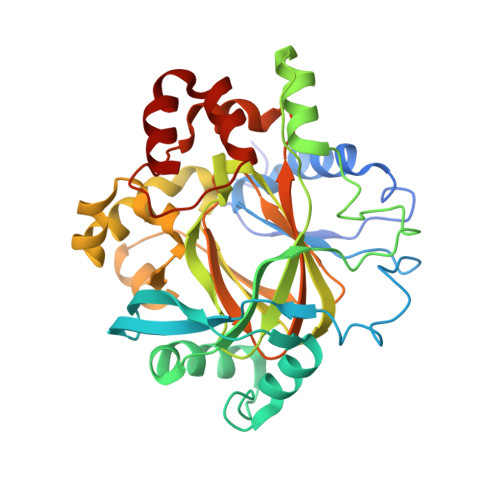
Structure-Based Screening of Tetrazolylhydrazide Inhibitors versus KDM4 Histone Demethylases.
Malecki, P.H., Ruger, N., Roatsch, M., Krylova, O., Link, A., Jung, M., Heinemann, U., Weiss, M.S.(2019) ChemMedChem 14: 1828-1839
- PubMed: 31475772 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/cmdc.201900441
- Primary Citation Related Structures:
6ETG, 6ETS, 6ETT, 6ETU, 6ETV, 6ETW - PubMed Abstract:
Human histone demethylases are known to play an important role in the development of several tumor types. Consequently, they have emerged as important medical targets for the treatment of human cancer. Herein, structural studies on tetrazolylhydrazide inhibitors as a new scaffold for a certain class of histone demethylases, the JmjC proteins, are reported. A series of compounds are structurally described and their respective binding modes to the KDM4D protein, which serves as a high-resolution model to represent the KDM4 subfamily in crystallographic studies, are examined. Similar to previously reported inhibitors, the compounds described herein are competitors for the natural KDM4 cofactor, 2-oxoglutarate. The tetrazolylhydrazide scaffold fills an important gap in KDM4 inhibition and newly described, detailed interactions of inhibitor moieties pave the way to the development of compounds with high target-binding affinity and increased membrane permeability, at the same time.
- Macromolecular Crystallography, Helmholtz-Zentrum Berlin für Materialien und Energie, Albert-Einstein-Str. 15, 12489, Berlin, Germany.
Organizational Affiliation: