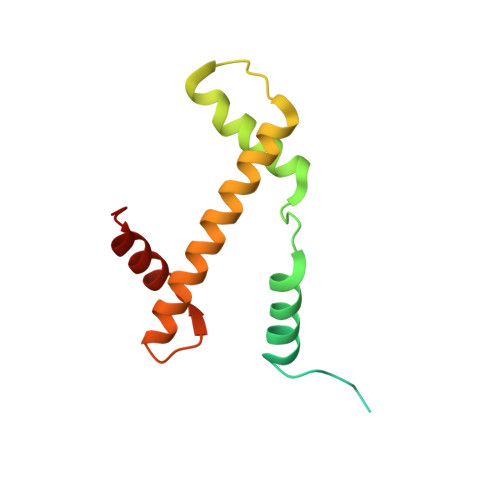
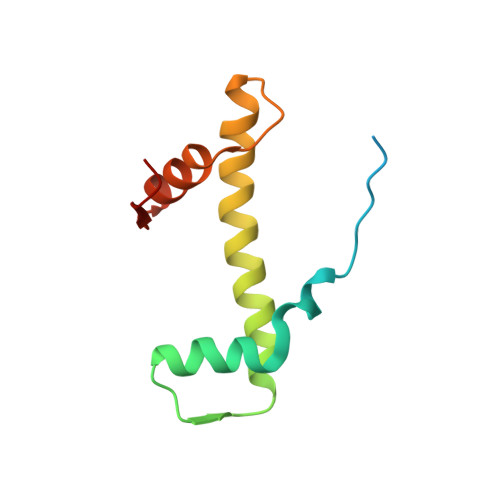
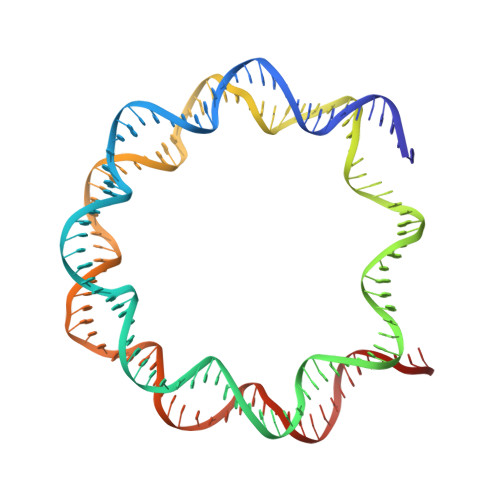
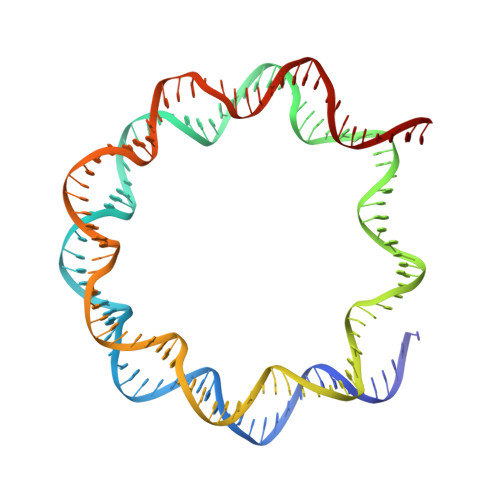
Histone octamer rearranges to adapt to DNA unwrapping.
Bilokapic, S., Strauss, M., Halic, M.(2018) Nat Struct Mol Biol 25: 101-108
- PubMed: 29323273 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-017-0005-5
- Primary Citation Related Structures:
6ESF, 6ESG, 6ESH, 6ESI - PubMed Abstract:
Nucleosomes, the basic units of chromatin, package and regulate expression of eukaryotic genomes. Although the structure of the intact nucleosome is well characterized, little is known about structures of partially unwrapped, transient intermediates. In this study, we present nine cryo-EM structures of distinct conformations of nucleosome and subnucleosome particles. These structures show that initial DNA breathing induces conformational changes in the histone octamer, particularly in histone H3, that propagate through the nucleosome and prevent symmetrical DNA opening. Rearrangements in the H2A-H2B dimer strengthen interaction with the unwrapping DNA and promote nucleosome stability. In agreement with this, cross-linked H2A-H2B that cannot accommodate unwrapping of the DNA is not stably maintained in the nucleosome. H2A-H2B release and DNA unwrapping occur simultaneously, indicating that DNA is essential in stabilizing the dimer in the nucleosome. Our structures reveal intrinsic nucleosomal plasticity that is required for nucleosome stability and might be exploited by extrinsic protein factors.
- Department of Biochemistry, Gene Center, LMU Munich, Munich, Germany.
Organizational Affiliation: