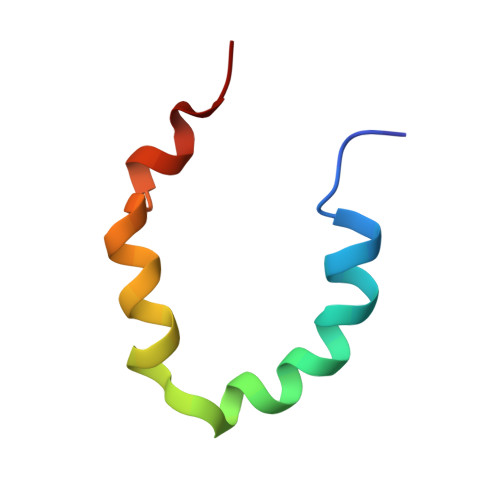
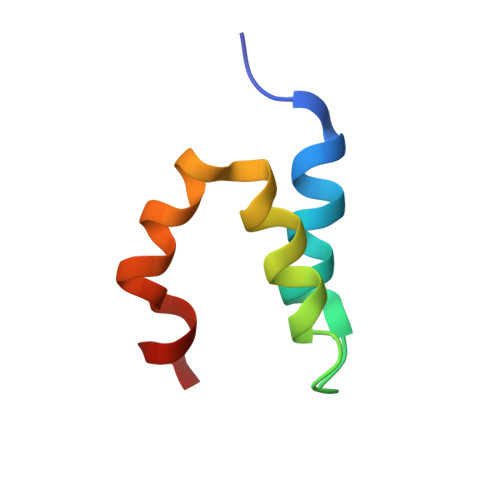
Structure and dynamics conspire in the evolution of affinity between intrinsically disordered proteins.
Jemth, P., Karlsson, E., Vogeli, B., Guzovsky, B., Andersson, E., Hultqvist, G., Dogan, J., Guntert, P., Riek, R., Chi, C.N.(2018) Sci Adv 4: eaau4130-eaau4130
- PubMed: 30397651 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.aau4130
- Primary Citation Related Structures:
6ES5, 6ES6, 6ES7 - PubMed Abstract:
In every established species, protein-protein interactions have evolved such that they are fit for purpose. However, the molecular details of the evolution of new protein-protein interactions are poorly understood. We have used nuclear magnetic resonance spectroscopy to investigate the changes in structure and dynamics during the evolution of a protein-protein interaction involving the intrinsically disordered CREBBP (CREB-binding protein) interaction domain (CID) and nuclear coactivator binding domain (NCBD) from the transcriptional coregulators NCOA (nuclear receptor coactivator) and CREBBP/p300, respectively. The most ancient low-affinity "Cambrian-like" [540 to 600 million years (Ma) ago] CID/NCBD complex contained less secondary structure and was more dynamic than the complexes from an evolutionarily younger "Ordovician-Silurian" fish ancestor (ca. 440 Ma ago) and extant human. The most ancient Cambrian-like CID/NCBD complex lacked one helix and several interdomain interactions, resulting in a larger solvent-accessible surface area. Furthermore, the most ancient complex had a high degree of millisecond-to-microsecond dynamics distributed along the entire sequences of both CID and NCBD. These motions were reduced in the Ordovician-Silurian CID/NCBD complex and further redistributed in the extant human CID/NCBD complex. Isothermal calorimetry experiments show that complex formation is enthalpically favorable and that affinity is modulated by a largely unfavorable entropic contribution to binding. Our data demonstrate how changes in structure and motion conspire to shape affinity during the evolution of a protein-protein complex and provide direct evidence for the role of structural, dynamic, and frustrational plasticity in the evolution of interactions between intrinsically disordered proteins.
- Department of Medical Biochemistry and Microbiology, Uppsala University, BMC Box 582, SE-75123 Uppsala, Sweden.
Organizational Affiliation: